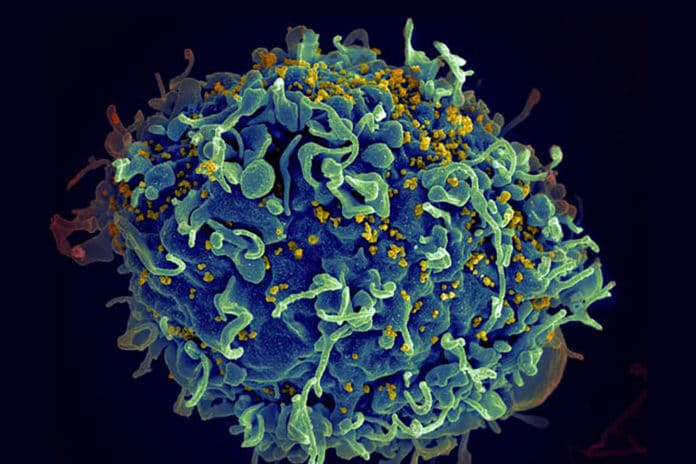
The adaptive immune system identifies and responds to different pathogens that humans experience throughout their lives. Deciphering immune cells’ ability to recognize harmful microbes is essential to fight off infection successfully. Although, the lack of technologies limits scientists’ efforts.
MIT biological engineers have developed a simple way to identify B or T cells that interact with viral or bacterial proteins to address this. They devised an experimental tool that discerns interactions between a particular immune cell and its target antigen.
Through this technique, scientists could understand how the immunity system itself works. It will help them make sense of complex immune recognition in various diseases and could accelerate the development of more effective vaccines and immunotherapies.
There are two main types of lymphocytes: B cells and T cells. The B cells produce antibodies for attacking invading bacteria, viruses, and toxins. The T cells destroy the infected or cancerous cells.
While encountering its target, T cells begin to proliferate to generate many identical cells to attack infected cells. On the other hand, B cells produce antibodies that help pick other components of the immune system to clear the infection.
Using tools, scientists identify specific antigen-immune cell interactions. However, these tools only allow identifying antigens exposed to one B or T cell or a large pool of immune cells encountering a small number of antigens.
The new tool developed by scientists allows the identification of huge libraries of both antigens and immune cells simultaneously. Scientists could pick out particular interactions within the vast realm of possibilities.
To develop this tool, scientists engineered a special form of lentivirus. Scientists generally use this type of virus to deliver genes as it can merge pieces of DNA into host cells. Lentivirus has an envelope protein called VSV-G that binds to receptors on the surface of many human cells, including immune cells, and infects them.
Scientists modified the VSV-G to avoid infection of cells. Instead, they used an antigen for the task. This modified version of VSV-G can only help the lentivirus enter cells if the paired antigen binds to a human B or T-cell receptor that recognizes the antigen.
Once the virus has entered, it integrates itself into the host cell’s genome. Therefore, by sequencing the genomes of all cells in the sample, scientists can find both the antigen expressed by the virus infecting the cell and the T or B-cell receptor sequence that allowed it to enter.
This way, scientists can use a viral infection itself to match up and then identify antigen-immune cell parings.
Scientists determined the tool’s efficiency by creating and screening different types of viruses against about 400,000 T cells. The tool could correctly pick out antigen-T-cell receptor pairings that had been previously identified.
Scientists also screened two different B-cell receptors against 43 antigens.
Michael Birnbaum is an associate professor of biological engineering at MIT, a member of MIT’s Koch Institute for Integrative Cancer Research, and the study’s senior author. He said, “The tool could accelerate the development of more effective vaccines and immunotherapies.”
“We built this tool to look for antigens, but there’s nothing particularly special about antigens. You could potentially use it to go into specific cells to do cell and gene therapy.”
Journal Reference:
- Dobson, C.S., Reich, A.N., Gaglione, S., et al. Antigen identification and high-throughput interaction mapping by reprogramming viral entry. Nat Methods (2022). DOI: 10.1038/s41592-022-01436-z