Steven Vokes, associate professor of molecular biosciences at the University of Texas at Austin, who became interested in Hedgehog, late in graduate school, while studying vascular development. He knew that Hedgehog signaling, a signaling pathway that transmits information to embryonic cells required for proper cell differentiation, is essential for vascular development as well as many other biological contexts.
It has been widely believed that proper embryonic development depends in large part on transcriptional repressors, proteins that bind to specific sites on DNA and prevent transcription of nearby genes. Transcriptional repression needs to be rapidly reversible during embryonic development. This extends to the Hedgehog pathway, which primarily serves to counter GLI (glioma-associated oncogene) repression by processing GLI proteins into transcriptional activators.
Speaking to Tech Explorist, Vokes said, “Although GLI proteins have now been studied for more than two decades, the mechanism by which they prevent target gene expression has remained unknown.”
While figuring out how GLI proteins work as transcriptional repressors, Vokes, along with his team found that the GLI repressors that keep the Hedgehog pathway turned off until the right time work by regulating chromatin on the level of enhancers, the essential DNA sequences that act as hubs for activating gene expression. The Hedgehog pathway does this by regulating biochemical modifications on chromatin-associated proteins. These chromatin modifications inactivate enhancers, which in turn prevent Hedgehog-regulated genes from being activated at the wrong time.
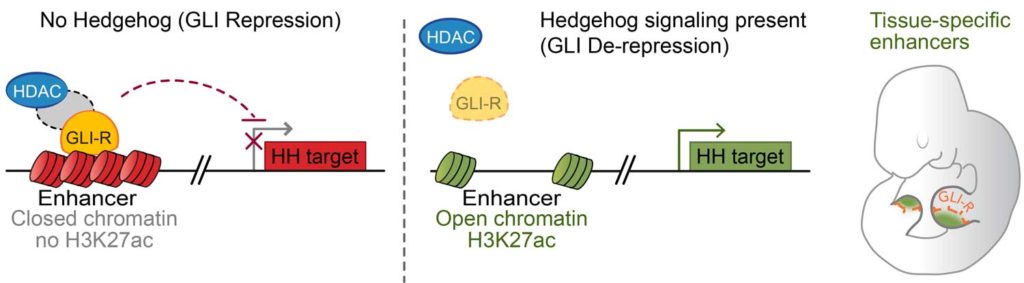
While explaining their finding to Tech Explorist, Vokes said, “GLI proteins repress transcription by binding to their enhancers, and I like to use an analogy to electrical wiring. You can regulate whether a light bulb is off or on (transcriptional output) by controlling the light switch, which is direct control. Or, if you are GLI proteins, they are controlling this upstream by regulating the activity of the electrical circuit that provides electricity to the switch.”
“This pathway is important from before the cradle to the grave, and it crosses so many different fields. It’s one of the keys to the body’s castle. The pathway itself is pretty well known, but the mechanisms at the DNA level are not.”
He further added, “The biggest challenge for the study was that GLI proteins function both as transcriptional activators (when Hedgehog signaling is around) and then are processed into transcriptional repressors when they are not. This bipotential role as an activator or repressor means that to understand the role of a repressor, you have to use double mutants that get rid of all GLI activity. Compounding the challenge of doing genomics in mutant backgrounds, we also were working with tiny embryonic tissues and had to optimize many protocols for small cell numbers.”
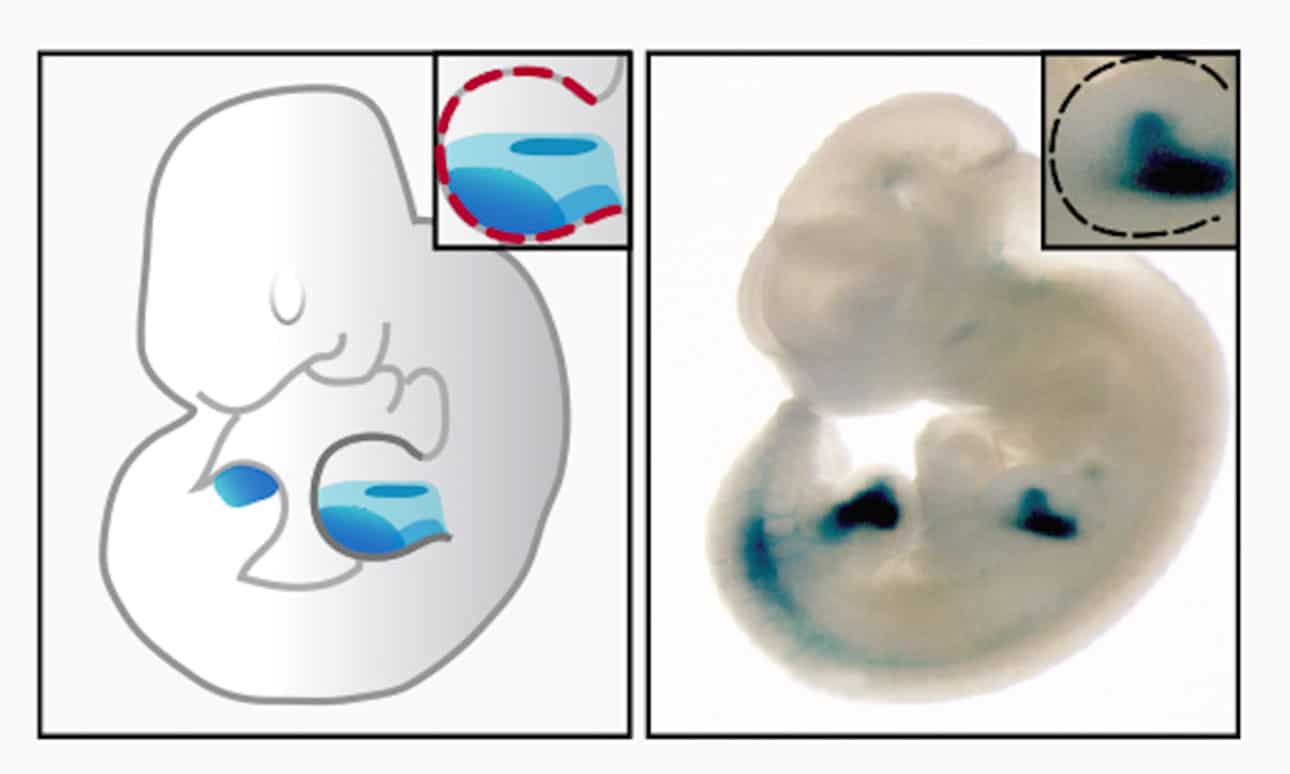
For their study, scientists used a genomic approach and the developing limb as a model and determined if GLI proteins repress Hedgehog target genes through altering the chromatin environment at GLI binding regions (GBRs). They found that GLI repressors regulate gene expression by inactivating enhancers.
A subset of GLI binding regions, termed Hedgehog-responsive enhancers, specifically loses acetylation in the absence of Hedgehog signaling. These regions are highly enriched around Hedgehog target genes and primarily drive Hedgehog-specific transcriptional activity in the mouse limb bud.
Also, scientists found that GLI repression regulates enhancer modification status, and thus, activity through the de-acetylation of Histone H3K27. This repression occurs independently of Polycomb activity. Enhancers regulated in this manner correspond to known GLI limb enhancers are highly enriched around Hedgehog target genes, and primarily drive tissue-specific enhancer activity within Hedgehog specific expression domains.
Highlighting the importance of the discovery, Vokes said, “First, that we now have an idea about how GLI repressors are controlling gene expression. This is important for literally dozens of different developmental pathways and enables us to start thinking about how we could control this activity. Second, we can now predict GLI enhancers that are likely to be actively regulating target gene expression based on their epigenetic response in mutant scenarios, something that was not possible to do before this research.”
Through this discovery, scientists might be able to unlock information on the genes casing specific congenital disabilities and offer better approaches to target cancers in a particular manner.
Other co-authors of the study include Rachel Lex, Kristen Falkenstein, and Joanna Henry of the University of Texas at Austin, as well as Zhicheng Ji, Weiqiang Zhou, and Hongkai Ji of Johns Hopkins University. Lex, Zhicheng Ji, and Falkenstein were co-first authors on the paper.
The study is published in the journal eLife.