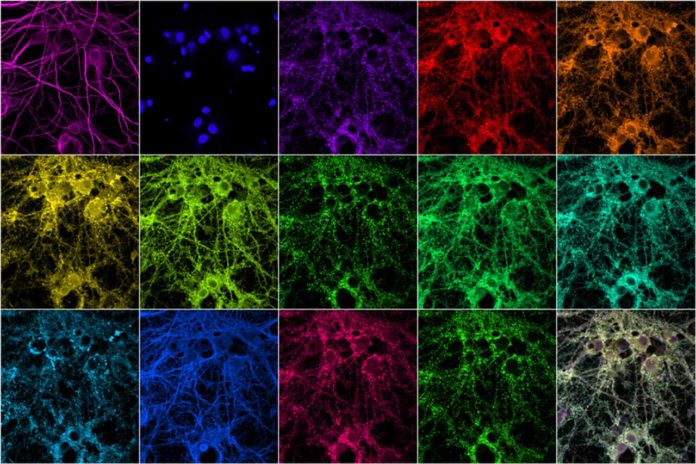
Scientists from MIT in collaboration with Broad Institute of Harvard have demonstrated a way to image these synaptic proteins at high resolution quickly.
Scientists used fluorescent nucleic acid probes to label and image an unlimited number of different proteins. They used this technique to label 12 different proteins found in the synapse, including scaffolding proteins, proteins associated with the cytoskeleton, and proteins that are known to mark excitatory or inhibitory synapses.
They developed this technique based on an existing method called DNA PAINT. The technique enables scientists to image protein by delivering a fluorescent DNA “oligo” that binds to the DNA-antibody probes.
The DNA strands have an inherently low affinity for one another, so they bind and unbind periodically, making a blinking fluorescence can be imaged utilizing super-resolution microscopy. Be that as it may, imaging each protein takes about thirty minutes, making it unfeasible for imaging numerous proteins in a large sample.
Scientists set out to make a quicker strategy that would enable them to dissect countless samples in a brief timeframe. To accomplish that, they changed the DNA-dye imaging probe, so it binds tie all the more tightly to the DNA-antibody, utilizing what is called locked nucleic acids. This gives a much brighter sign, so the imaging should be possible all the more rapidly, yet at a somewhat lower resolution.
Mark Bathe, an MIT associate professor of biological engineering said, “Inhibitory and excitatory are the canonical synapse types, but it is speculated that there are numerous different subtypes of synapses, without any real consensus around what those are.”
Scientists also show that they could measure changes in synaptic protein levels that occur after neurons are treated with a compound called tetrodotoxin (TTX), which strengthens synaptic connections.
Eric Danielson, an MIT senior postdoc who is an author of the study said, “Using conventional immunofluorescence, you can typically extract information from three or four targets within the same sample, but with our technique, we were able to expand that number to 12 different targets within the same sample. We applied this method to examine synaptic remodeling that occurs following treatment with TTX, and our finding corroborated previous work that revealed a coordinated upregulation of synaptic proteins following TTX treatment.”
Scientists are now using this technique, called PRISM, to study how the structure and composition of synapses are affected by knocking down a set of genes reported previously to confer genetic risk for development of psychiatric disorders.
Karen Perez de Arce, a Broad Institute research scientist and an author of the study said, “With this approach, we expect to provide a more detailed overview of the changes in synaptic organization and shared disease effects associated with these genes.”
Bathe said, “Understanding how genetic variation impacts neurons’ development in the brain, and their synaptic structure and function, is a huge challenge in neuroscience and in understanding how these diseases arise.”
The study is published in the journal Nature Communications. The National Institutes of Health-funded the research, including the NIH BRAIN Initiative, the National Science Foundation, the Howard Hughes Medical Institute Simons Faculty Scholars Program, the Open Philanthropy Project, the U.S. Army Research Laboratory, the New York Stem Cell Foundation Robertson Award, and the Stanley Center for Psychiatric Research.
Other authors of the paper include MIT research scientist Demian Park, former MIT graduate student Anthony Kulesa, and MIT postdoc Eike-Christian Wamhoff. Paul Blainey, an associate professor of biological engineering and a member of the Broad Institute, and Edward Boyden, the Y. Eva Tan Professor in Neurotechnology and an associate professor of biological engineering and brain and cognitive sciences are also authors of the study.