The study investigates a fascinating connection between two seemingly unrelated phenomena: sperm swimming and the formation of zebra stripes. Researchers from the University of Bristol have uncovered that the patterns governing the motility of sperm are strikingly similar to the ways that dictate the distinctive stripes on zebra coats.
Today’s study, published in Nature Communications, uncovers a surprising connection. It turns out that the way sperm tails and tiny hair-like structures called cilia move follows a pattern discovered by the famous mathematician Alan Turing.
These moving structures create stripe-like patterns over time and space, generating waves that propel sperm and microorganisms forward.
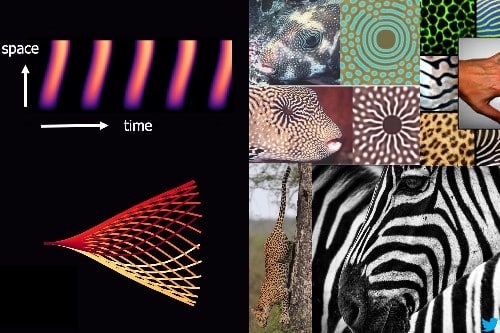

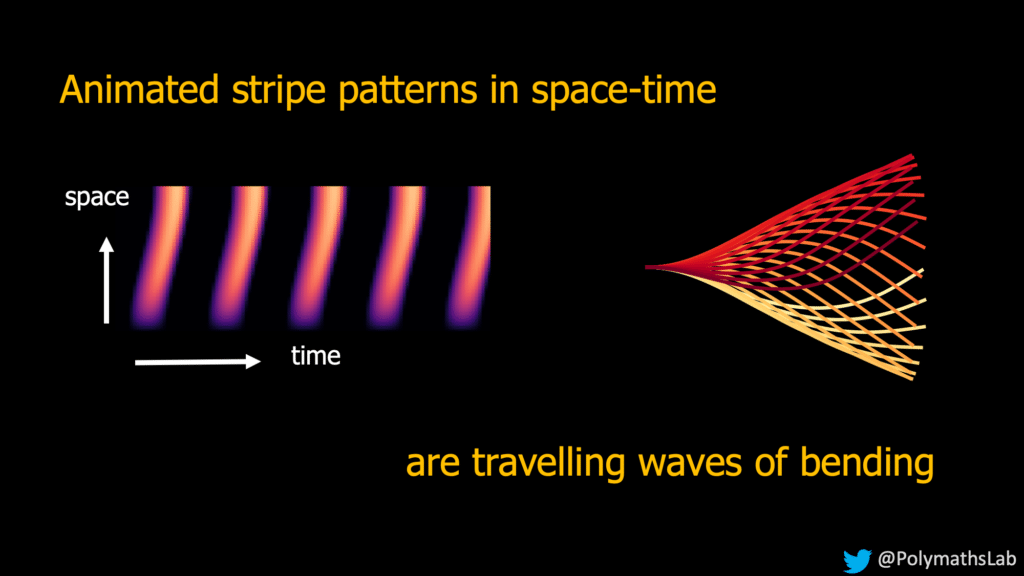
You might know Alan Turing for his role in breaking the enigma code during World War II. However, he also developed a theory about how patterns can form. He said that “marks can appear independently with just two things: chemicals spreading out and reacting together. This idea is called the reaction-diffusion theory for pattern formation, and it’s the same principle at work in the movement of sperm tails and cilia.”
Turing pioneered reaction-diffusion math to study natural patterns, leading to what we now call Turing patterns. While not yet confirmed by experiments, these patterns are believed to influence diverse natural designs like leopard spots, sunflower seed arrangements, and beach sand patterns. Turing’s theory has broad applications, from biology and robotics to astrophysics, shaping various fields of inquiry.
Mathematician Dr. Hermes Gadêlha, head of the Polymaths Lab, and his Ph.D. student James Cass conducted this research in the School of Engineering Mathematics and Technology at the University of Bristol. Gadêlha explained: “Live spontaneous motion of flagella and cilia is observed everywhere in nature, but little is known about how they are orchestrated.“
“They are critical for the health and disease, reproduction, evolution, and survival of almost every aquatic microorganism on earth,” he added.
The team was inspired by recent findings in thin fluids that suggested the environment doesn’t affect flagellum much. They used math, simulations, and data to prove that flagella can move independently without needing their fluid surroundings.
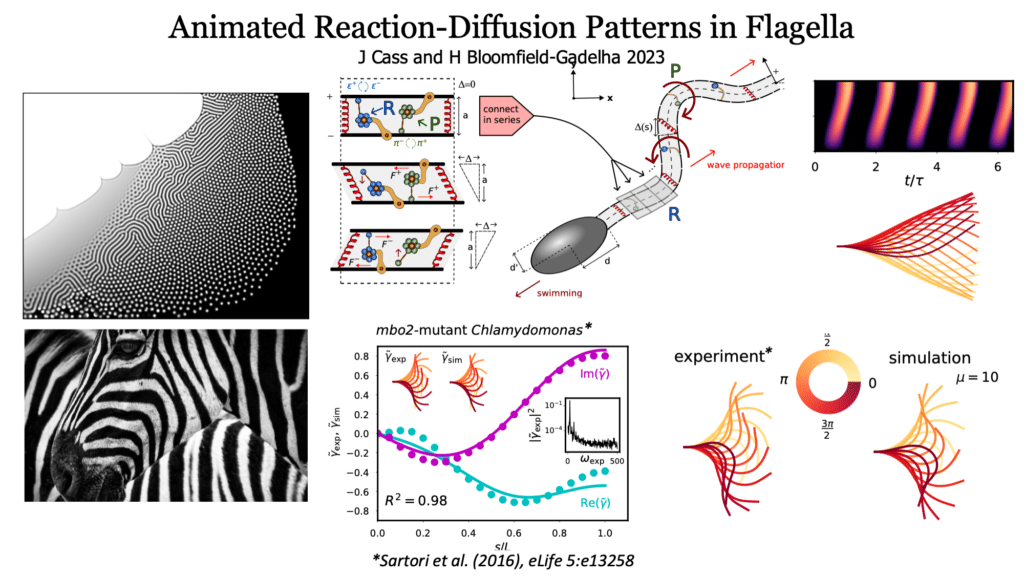
This is similar to what Alan Turing talked about with chemical patterns. In the case of sperm, molecular motors cause chemical reactions, powering the flagellum. The bending motion then spreads like waves along the tail. It’s surprising how similar this is to how patterns look, and it shows that you only need two simple things to make very complex movements happen.
Dr. Gadêlha mentioned that they found a mathematical “recipe” that both bull sperm and a type of green algae called Chlamydomonas follow. This suggests that nature uses similar solutions in different species.
These swimming waves in the flagellum happen even when not influenced by the fluid around it. This is a reliable way for aquatic species to swim in thin liquids.
Thanks to the researchers who shared their data, the study’s simulations matched real-world data for the first time. These findings could help us understand fertility issues linked to flagellar problems and ciliopathies, which are diseases caused by ineffective cilia in human bodies, in the future.
This discovery has potential uses in robotics, artificial muscles, and animated materials. The team found a simple “mathematical recipe” for creating movement patterns.
Dr. Gadêlha is part of the SoftLab at Bristol Robotics Laboratory, where he uses math to develop advanced soft robots.
He explains that in 1952, Turing unlocked the secret behind chemical patterns. They’ve shown that tiny cellular structures, like flagella, use Turing’s idea to create motion designs, like sperm tail movement.
However, this model is still relatively basic, and there might be more complex ones we have yet to learn about. So, more research is needed. The Engineering and Physical Sciences Research Council (EPSRC) funded the study. It was supported by a DTP studentship for James Cass’s PhD.
The numerical work was done using the University of Bristol’s Advanced Computing Research Centre’s facilities.
Journal Reference:
- Cass, J.F., Bloomfield-Gadêlha, H. The reaction-diffusion basis of animated patterns in eukaryotic flagella. Nature Communications. DOI: 10.1038/s41467-023-40338-2.