A team of researchers led by Baylor College of Medicine and the Czech Academy of Sciences has revealed a new piece of the puzzle of how gene expression is arranged. A study published in the journal Science showed findings that will reveal the mechanism which coordinates the assembly of components inside cells that control gene expression. The mechanism will be essential for normal cell function. It will also have implications for the treatment of cancer, neurodegeneration, and HIV infection.
“Most previous studies have focused on particular cellular components that turn genes completely on or off,” said co-corresponding author Dr. H. Courtney Hodges, assistant professor of molecular and cellular biology and in the Center for Precision Environmental Health at Baylor. “Our work reveals a new perspective – that the proteins that regulate the rate of gene expression also can work collectively to finely tune expression levels in many different settings. We identified a mechanism that brings these proteins together and plays widespread roles in health and disease.”
In this study, the researchers extended the study of TNDs (TFIIS N-terminal domains) and found them in many other proteins.
“Everywhere we looked, we found these domains, in particular in the machinery that regulates transcription elongation, one of the first steps of gene expression in all human cells. Transcription elongation is a complex cellular process that involves many different proteins working together,” said first author Dr. Katerina Cermakova, a postdoctoral fellow in the Hodges lab. “We discovered that TNDs are the most enriched structural element in all transcription elongation factors. Once you look for them, you find that all the important protein complexes involved in transcription elongation have a TND or bind a protein that has one.”
“One remarkable thing about these unstructured regions is their unusual behavior as molecules,” said co-corresponding author Dr. Vaclav Veverka, a structural biologist and group leader at the Institute of Organic Chemistry and Biochemistry of the Czech Academy of Sciences (IOCB Prague). “Imagine a TIM as a string that is loose at one end and moves as if being blown around in a hurricane. But when it finds its TND partner, the string curls up and holds on very tightly to the TND to keep it close.” The researchers show this attachment plays an important role in the early stages of gene expression.
“We first determined that TNDs and TIMs bound together in ‘test tube’ type of experiments, but it was really exciting to see that they bind to each other in living cells, validating the relevance of our observations in living systems,” Cermakova said. “We also determined that the TND-TIM interactions are highly specific.”
“I was surprised to see that IWS1, a protein previously thought to be a secondary player in the transcription elongation machinery, acts as a central organizer of these factors,” said Hodges, a member of Baylor’s Dan L Duncan Comprehensive Cancer Center.
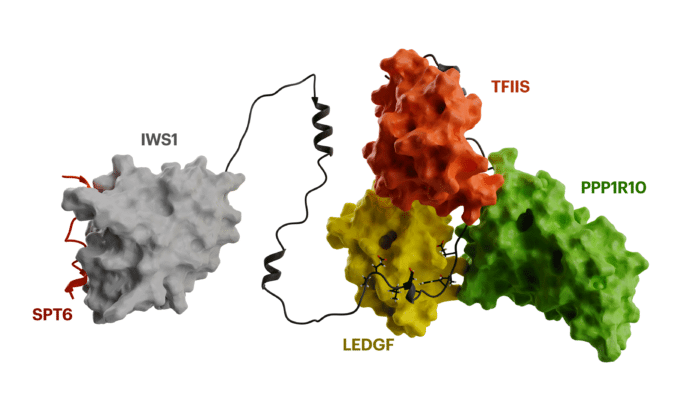
“We found that IWS1 uses specific TND-TIM interactions to coordinate the activities of many transcription regulators at the same time, making it appear like a conductor at a symphony that keeps all the factors working in harmony and close by,” Veverka said.
The team also explored the consequences of disrupting a single unstructured protein region on the harmony of the transcription elongation process.
“Hundreds of genes with important functions were altered when we disrupted even a single unstructured region,” Hodges said. “The first step of gene expression started, but was paused and unable to be completed, preventing efficient gene expression.”
The study highlights the uncredited role of disordered protein interactions as key arrangers in gene expression and other complex biological functions. The findings also can contribute to a better understanding of diseases such as cancer, viral infections, neurodevelopmental disorders, and potentially other conditions in which these factors are disrupted. TNDs and TIMs may represent essential novel targets for improved treatments for these conditions.
Journal Reference
- Kateřina Čermáková, Jonas Demeulemeester, Vanda Lux, Monika Nedomová, Seth R. Goldman, Eric A. Smith, Pavel Srb, Rozálie Hexnerová, Milan Fábry, Marcela Mádlíková, Magdalena Hořejší. Jan De Rijck, Zeger Debyser, Karen Adelman, H. Courtney Hodges, Václav Veverka. A ubiquitous disordered protein interaction module orchestrates transcription elongation. Science 2021. DOI: 10.1126/science.abe2913