Scientists at Brown University have taken a step forward toward storing huge amounts of data in molecular storage systems. The study has shown that it’s possible to store and retrieve data stored in artificial metabolomes — arrays of liquid mixtures containing sugars, amino acids, and other types of small molecules.
As a proof of principle of small-molecule postgenomic data storage, scientists demonstrate a workflow for representing abstract data in synthetic mixtures of metabolites. They have encoded kilobyte-scale image files into metabolite solutions and read the information back out again.
Jacob Rosenstein, a professor in Brown’s School of Engineering and senior author of the study said, “This is a proof-of-concept that we hope makes people think about using wider ranges of molecules to store information. In some situations, small molecules like the ones we used here can have even greater information density than DNA.”
“Another potential advantage stems from the fact that many metabolites can react with each other to form new compounds. That creates the potential for molecular systems that not only store data, but also manipulate it — performing computations within metabolite mixtures.”
A key strength of this work is that it can be applied to any chemical library. Metabolites hold particular potential because they provide access to well-regulated interconversion networks, materials, and databases which could facilitate computational operations on chemical data.
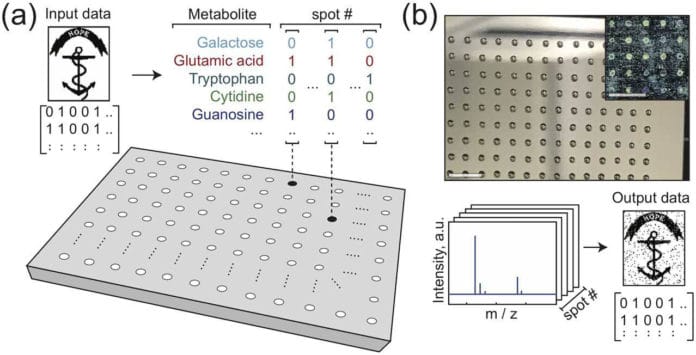
The presence or absence of one metabolite in one spot encodes one bit of information. Therefore, the total number of bits stored in one place is equal to the number of available library elements.
Scientists wanted to check if artificial metabolomes could store data. For this, they assembled their artificial metabolomes — small liquid mixtures with different combinations of molecules. The presence or absence of a particular metabolite in a mix encodes one bit of digital data, a zero or a one.
Researchers assembled their very own counterfeit metabolomes — small liquid mixtures with various combination of molecules. The presence or absence of a specific metabolite in a mix encodes one bit of digital data, a zero or a one. The quantity of molecule types in the artificial metabolome decides the number of bits every mixture can hold.
For this investigation, the analysts made libraries of six and 12 metabolites, which means every mixture could encode either six or 12 bits. A large number of mixtures are then arrayed on small metal plates as nanoliter-sized droplets. The contents and arrangement of the droplets, correctly put by a fluid taking care of robot, encodes the ideal information.
The plates are then dried, leaving minor spots of metabolite atoms, each holding digital data. The data would then be able to be read out utilizing a mass spectrometer, which can distinguish the metabolites present at each spot on the plate and decode the data.
Using the technique, scientists were able to tp encode and retrieve a variety of image files of sizes up to 2 kilobytes.
Scientists noted, “That’s not big compared to the capacity of modern storage systems, but it’s a solid proof-of-concept. And there’s plenty of potential for scaling up. The number of bits in a mixture increases with the number of metabolites in an artificial metabolome, and there are thousands of known metabolites available for use.”
“Currently, there are some limitations. For example, many metabolites chemically interact with each other when placed in the same solution, and that could result in errors or loss of data. But that’s a bug that could ultimately become a feature. It may be possible to harness those reactions to manipulate data — performing in-solution computations.”
Brenda Rubenstein, a Brown assistant professor of chemistry and co-author of the study, said, “Using molecules for computation is a tremendous opportunity, and we are only starting to figure out how to take advantage of it. Research like this challenges what people see as being possible in molecular data systems. DNA is not the only molecule that can be used to store and process information. It’s exciting to recognize that there are other possibilities out there with great potential.”
Other authors on the paper are Christopher Arcadia, Joseph Geiser, Peter Weber, and Christopher Rose. The research was supported by DARPA (W911NF-18-2-0031).