Molecular engineering aims to develop modular functional units that can be used in the bottom-up design of nanoassemblies capable of carrying out challenging tasks. These systems need fuel-consuming nanomotors that can actively push passive followers downstream. With rare exceptions, Brownian motion drives the majority of artificial molecular motors, but the forces that are created are typically non-directed and insufficient to effectively transmit to passive second-level components. As a result, effective nanoscale driver-follower systems powered by chemical fuel have yet to be developed.
A new study by an international team of scientists presented a DNA nanomachine- a novel type of nano-engine made of DNA. It can make pulsing movements and is propelled by an intelligent mechanism. The idea is for the researchers to install it as a drive in intricate nanomachines after fitting it with a connection.
Scientists used the group’s computer modeling tools to gain insights into the design and operation of this leaf-spring nano-engine. Nearly 14,000 nucleotides, DNA’s fundamental structural building blocks, make up the structure.
Petr Šulc, an assistant professor at Arizona State University‘s School of Molecular Sciences and the Biodesign Center for Molecular Design and Biomimetics, said, “Being able to simulate motion in such a large nanostructure would be impossible without oxDNA, the computer model that our group uses for design and design of DNA nanostructures.”
“It is the first time a chemically powered DNA nanotechnology motor has been successfully engineered. We are very excited that our research methods could help with studying it, and are looking forward to building even more complex nanodevices in the future.”
This innovative engine functions similarly to a grip-strengthening hand exerciser when used frequently. The motor, however, is around a million times smaller. In a V-shape, two handles are joined by a spring.
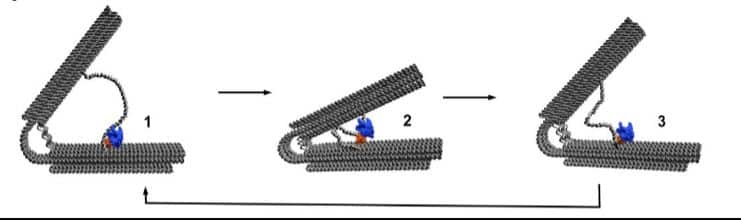
Professor Michael Famulok (project lead) from the University of Bonn, Germany, said, “In a hand grip strength trainer, you squeeze the handles together against the resistance of the spring. Once you release your grip, the spring pushes the handles back to their original position. Our motor uses a very similar principle. But the handles are not pressed together but rather pulled together.”
According to the experts, without a specific mechanism, there wouldn’t be any plants or creatures on Earth. There is a small library in every cell. It contains the instructions for every kind of protein a cell needs to function. The cell requests a copy from the appropriate blueprint to manufacture a specific type of protein. The RNA polymerases are the enzymes responsible for creating this transcript.
Long DNA strands make up the initial blueprint. These strands are followed by the RNA polymerases, which copy the information stored there letter by letter.
Famulok said, “We took an RNA polymerase and attached it to one of the handles in our nanomachine. Nearby, we also strained a DNA strand between the two handles. The polymerase grabs onto this strand to copy it. It pulls along the strand, and the non-transcribed section becomes increasingly smaller. This pulls the second handle bit by bit towards the first one, compressing the spring simultaneously.”
Just before it ends, the DNA strand between the handles carries a specific set of letters. This so-called termination sequence instructs the polymerase to release the DNA from its grasp. Once more relaxed, the spring can then move the handles apart. As a result, the molecular copier can initiate a new transcription process because the start sequence of the strand is now near the polymerase. The process then continues.
Mathias Centola, who is part of the research group headed by Famulok and carried out a large proportion of the experiments, said, “In this way, our nanomotor performs a pulsing action.”
Like all motors, this one requires energy to operate. It is given by the “alphabet soup” from which the polymerase makes the transcripts. Each of these letters, or nucleotides in scientific jargon, has a little tail of three phosphate groups, or a triphosphate. The polymerase must eliminate two phosphate groups to add a new letter to an existing phrase. As a result, energy is released that can be used to connect the letters.
Famulok said, “Our motor thus uses nucleotide triphosphates as fuel. It can only continue to run when a sufficient number of them are available.”
Scientists next demonstrated that the motor can be combined with other structures. It should be able to do things like meander across a surface, much like an inchworm that pushes itself along a branch uniquely.
Famulok said, “We are also planning to produce a type of clutch that will allow us only to utilize the motor’s power at certain times and otherwise leave it to idle. In the long term, the motor could become the heart of a complex nanomachine. However, much work must be done before we reach this stage.”
In particular, the group develops new multiscale models for creating and modeling DNA and RNA nanostructures and devices to examine interactions between biomolecules.
Šulc said, “Just as complex machines in our everyday use — planes, cars, and chips in electronics — require sophisticated computer-aided design tools to make sure they perform a desired function, there is a pressing need to have access to such methods in the molecular sciences.”
Journal Reference:
- Centola, M., Poppleton, E., Ray, S. et al. A rhythmically pulsing leaf-spring DNA-origami nanoengine that drives a passive follower. Nat. Nanotechnol. (2023). DOI: 10.1038/s41565-023-01516-x
