Rice University researchers have viably extinguished a verbal confrontation over the component behind fluorescent biosensors that screens neurons by detecting changes in voltage.
The work drove by Rice hypothetical physicist Peter Rossky and postdoctoral scientist Lena Simine affirmed through PC reproductions their hypothesis that a mechanical procedure controls the extinguishing of fluorescence in ArcLight, an engineered voltage marker put inside proteins that line the inward films of neurons.
Through their models, the scientists coupled both the instrument and fluorescence to the quality of electric fields they saw over the chromophore, the fluorescing some portion of the protein. Their outcomes demonstrated a straightforward measure of the field in a recreation could be utilized to foresee whether and how well new fluorescent sensors will carry on before analysts blend them, Rossky said.
The examination shows up in the Journal of the American Chemical Society.
ArcLight, created by Yale neuroscientist Vincent Pieribone in 2012, is a hereditarily encoded fluorescence voltage pointer protein. It contains a transformation that influences the fluorescence to flag diminish when the voltage rises and light up when the voltage falls. That makes it valuable for following signs in the sensory system by communicating the protein in neurons and perceiving how they illuminate.
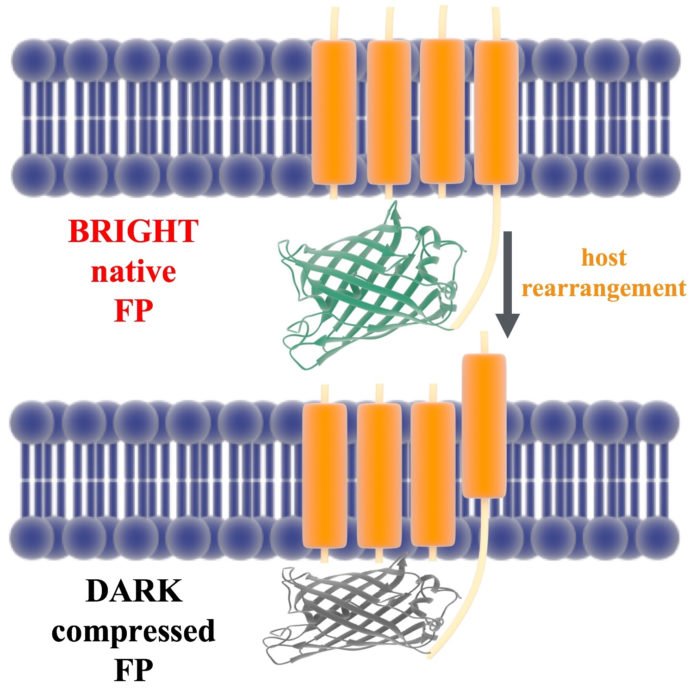
The protein is fastened to the neuron’s phone divider by a voltage-detecting segment that moves a couple of angstroms when a flag from another neuron changes the electrical charge in the film. The Rice analysts speculated that movement pulls the protein against the film, packing it and extinguishing fluorescence.
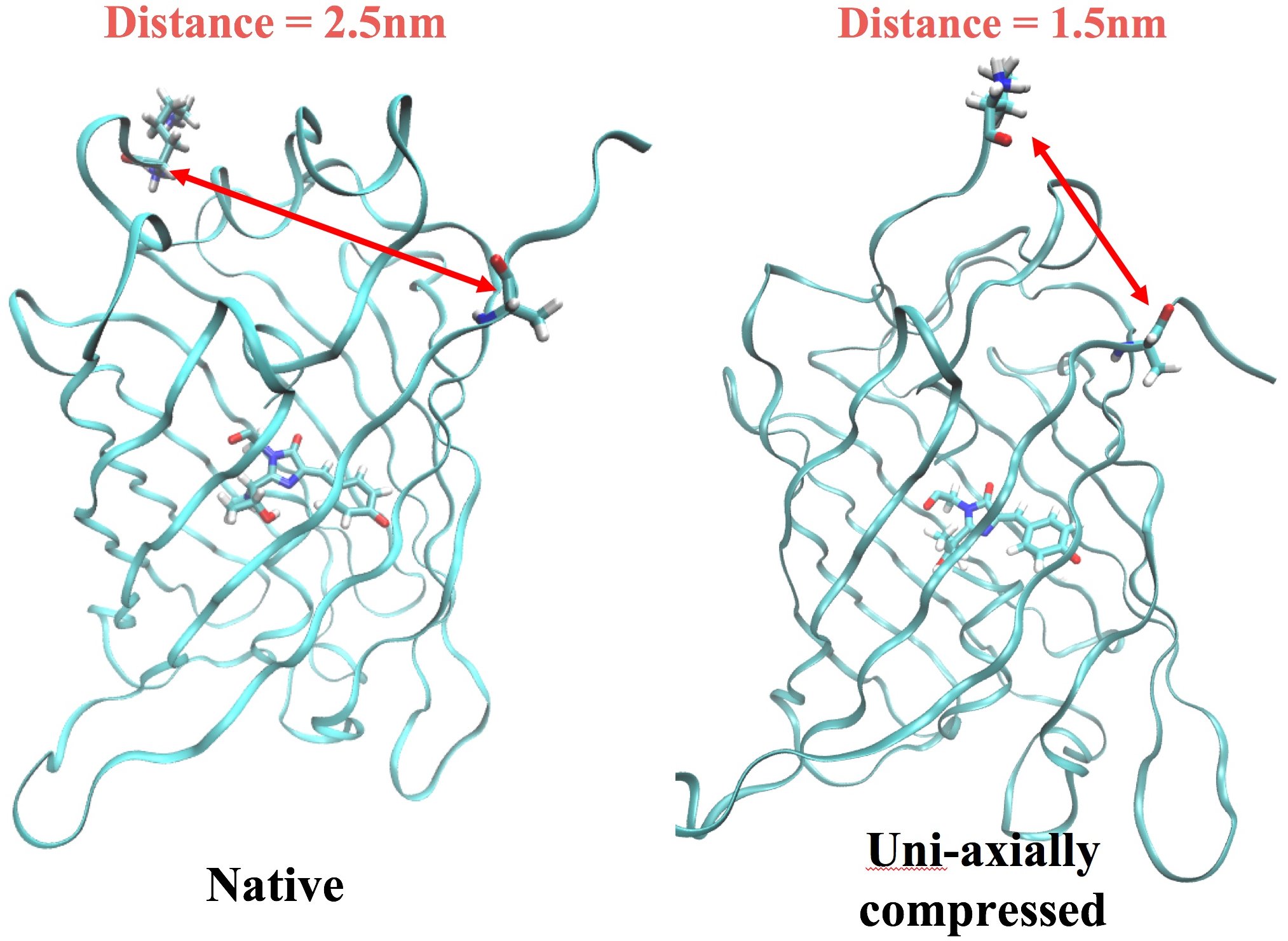
Rossky said changing the state of the protein conveys two deposits a nanometer nearer to each other. No more to manage how the chromophore disposes of vitality, either as light (by surrendering photons and fluorescing) or as warmth.
“We hypothesized what geometry change occurs in the protein as a result of the response of the membrane. ‘Does this change the fluorescence?’ And we found that it does. In addition, we showed that monitoring a much simpler quality — the electric field along two axes where the fluorescence comes from — is sufficient to completely describe the response.”
“I thought, ‘That sounds like a good project for me’.”
Working with analysts in the gathering of José Onuchic at Rice’s Center for Theoretical Biological Physics (CTBP) permitted Simine, a concoction physicist via preparing, to exploit the inside’s aptitude in reproducing proteins for testing.
She said a decade-long wrangle between researchers neglected to decide if mechanical or electrical properties of proteins caused their fluorescence. It ended up being a touch of both.
Simine said, “A recent paper gave computational evidence for it being predominantly electrostatic, and it kind of makes sense because the protein’s very soft. We also figured those mutations are sticking to the membrane, and when they do, the protein’s orientation allows the protein to be compressed.”
She found electrostatic changes to the neuronal membrane triggered the physical change that quenches fluorescence but also left an electrical trace in the protein that could be observed in the simulation.
“We put some thought into it and came up with a reaction coordinate. We can take any mutation of the sequence of this protein and translate it into two numbers that are the inputs for this model, the electrostatic fields around the chromophore. It’s a nice, elegant phenomenological theory.”
The lab intends to test its procedure on exclusively blended fluorescent proteins and coordinating recreations to check whether their hypothesis and experimentation keep on aligning. On the off chance that they do, they expect their models will be very valuable to manufactured scholars making new classes of fluorescent markers.