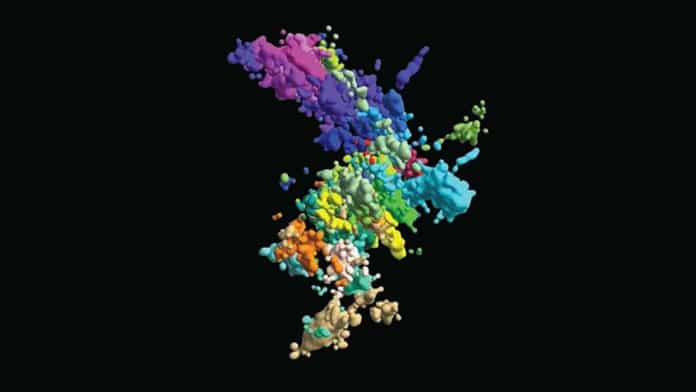
Cellular DNA always forms a complex with various protein partners that help package it into such a tiny space. This DNA-protein complex is called chromatin, wherein the mass of protein and nucleic acid is nearly equal.
Within cells, chromatin usually folds into characteristic formations called chromosomes. Each chromosome contains a single double-stranded piece of DNA along with the aforementioned packaging proteins.
When fully condensed, replicated chromosomes appear as thick X-shaped structures. This is how our high-school textbooks taught us. But, those images are far from accurate.
Jun-Han Su from Harvard University said, “For 90 percent of the time, chromosomes don’t exist like that.”
Last year, before Su graduated with his Ph.D., he and three current Ph.D. candidates in the Graduate School of Arts and Sciences—Pu Zheng, Seon Kinrot, and Bogdan Bintu—captured high-resolution 3-D images of human chromosomes, the complex houses for our DNA. Now, those images could provide enough evidence to change those Xs into more complex but far more accurate symbols to not only teach the next generation of scientists but help the current generation unravel mysteries about how chromosome structure influences function.
Cells divide and replicate their DNA within labyrinthine libraries inside chromatin, the stuff inside chromosomes.
Extended in a straight line, DNA in a single cell can reach six feet, all of which gets wrapped into tight, complex structures in a cell nucleus. Just one mistake copying or re-winding that genetic material could cause genes to mutate or malfunction.
In a new study, Zhuang and her team report a new method to image chromatin structure and behavior together. The method includes connecting the dots to determine how one influences the other to maintain proper function or cause disease.
Zhuang, David B. Arnold, Jr. Professor of Science, said, “It’s quite important to determine the 3-D organization to understand the molecular mechanisms underlying the organization and also to understand how this organization regulates genome function.”
Using a high-resolution 3-D imaging method, the team built a chromosomal map from both wide-lens images of all 46 chromosomes and close-ups of one section of one chromosome. They also captured connected dots (“genomic loci”) along each DNA chain to image something that’s still too small to image. Connecting lots of dots formed a comprehensive picture of the chromatin structure.
Zhuang said, “But there was a snag. Previously, the number of dots they could image and identify was limited by the number of colors they could image together: three. Three dots can’t make a comprehensive picture.”
Hence, the team came up with an approach that includes imaging three different loci, quenching the signal, and then imaging another three in rapid succession. With that technique, each dot gets two identifying marks: color and image round.
Zhuang said, “Now we have 60 loci simultaneously imaged and localized and, importantly, identified.”
Still, to cover the whole genome, they needed more—thousands—so they turned to a language that’s already used to organize and store huge amounts of information: binary. By imprinting binary barcodes on different chromatin loci, they could image far more loci and decode their identities later.
With 20-bit barcodes, the team could differentiate 2,000 molecules in just 20 rounds of imaging.
Using the technique, the team could image about 2,000 chromatin loci per cell- enough to form a high-resolution image of what the structure of chromosomes looks like in its native habitat. Additionally, they imaged transcription activity (when RNA replicates genetic material from DNA) and nuclear structures like nuclear speckles and nucleoli.
With their 3-D Google Maps of the genome, they could analyze how the structure shifts over time and how those territorial movements help or hurt cell division and replication.
It is well known that chromatin is broken into different areas and domains (like deserts versus cities). But how those terrains appear remains obscure.
Zhuang and team determined that areas with lots of genes (“gene-rich”) tend to flock to similar areas on any chromosome with their high-resolution images. But regions with few genes (“gene-poor”) only come together if they share the same chromosome.
Despite requiring more efforts, the study makes sure that the local chromatin environment impacts transcription activity. The structure does influence function.
What’s more, the team also discovered that two chromosomes look so different from each other. They do not look the same, even in otherwise identical cells. To find out what each chromosome seems like in every cell in the human body will take far more work than one lab can take on alone.
Zhuang said, “It’s not going to be possible to build just on our work. We need to build on many, many labs’ work to have a comprehensive understanding.”
Journal Reference:
- Jun-Han Su et al., Genome-Scale Imaging of the 3D Organization and Transcriptional Activity of Chromatin, Cell (2020). DOI: 10.1016/j.cell.2020.07.032