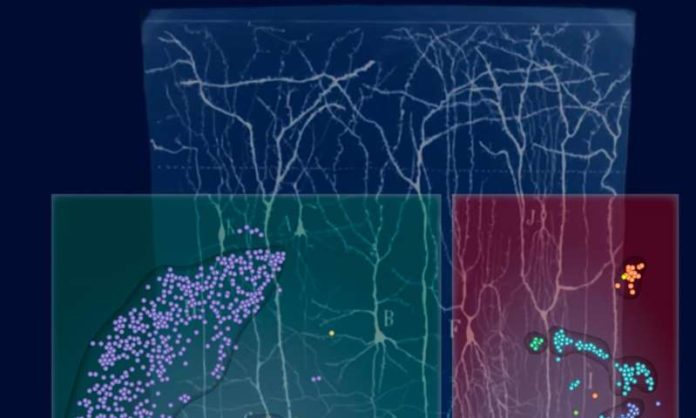
It is hard to identify the difference between two neurons or brains cells under the microscope. So researchers have discovered atomic strategies to distinguish gatherings of neurons with various capacities.
For the first time, scientists at Salk Institute and University of California San Diego profiled synthetic alterations of DNA atoms in singular neurons. This gives them detailed information about what makes brain cells different from its neighbor.
According to scientists, this could allow prompting a drastically better comprehension about mental health and brokenness.
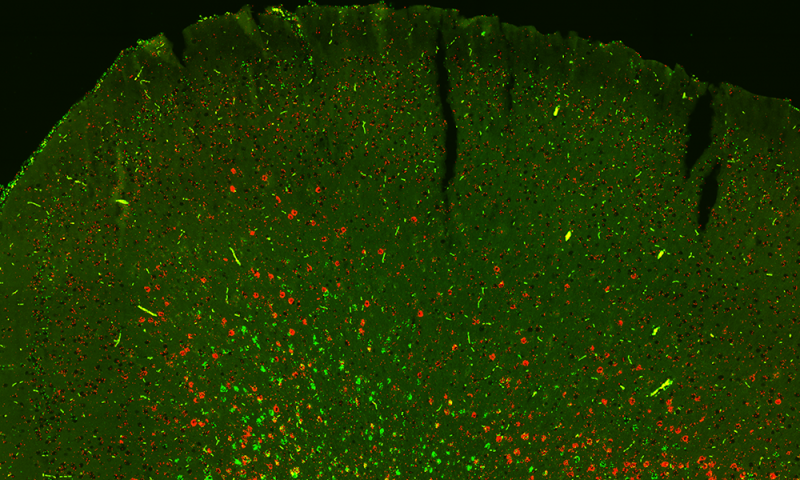
Scientists noted, “This is a critical step in beginning to identify how many types of neurons exist. Each cell’s methylome, the pattern of chemical markers made up of methyl groups that study its DNA gave a distinct readout that helped the Salk team sort neurons into subtypes.”
“We think it’s pretty striking that we can tease apart a brain into individual cells, sequence their methylomes, and identify many new cell types along with their gene regulatory elements, the genetic switches that make these neurons distinct from each other.”
Primarily, scientists were seemed as using RNA molecules inside individual brain cells. But there was a problem that RNA levels rapidly change when a cell is exposed to new conditions, or even throughout the day. So, instead of using RNA molecules, scientists used cells’ methylomes, which are generally stable throughout adulthood.
The study clearly defines neuronal types based on their methylomes. So, according to that, it could open up the possibility of understanding what makes two neurons that sit in the same brain region and otherwise look similar behave differently.
Scientists focused on both mouse and human brains by focusing on the frontal cortex. They disengaged 3,377 neurons from the frontal cortex of mice and 2,784 neurons from the frontal cortex of an expired 25-year-old human.
Scientists then used a method called snmC-seq to sequence the methylomes of each cell. As the neurons have two types of methylation, the approach mapped both types methylation- CG methylation and non-CG methylation.
They found that the neurons in mouse frontal cortex, clustered into 16 subtypes based on methylation patterns. On the other hand, the human frontal cortex was more diverse and formed 21 subtypes. Inhibitory neurons those that provide stop signals for messages in the brain showed more conserved methylation patterns between mice and humans compared to excitatory neurons.
Eran Mukamel of the UC San Diego Department of Cognitive Science said, “This study opens a new window into the incredible diversity of brain cells.”
Chongyuan Luo, a Salk research associate said, “There are hundreds, if not thousands, of types of brain cells that have different functions and behaviors and it’s important to know what all these types are to understand how the brain works. Our goal is to create a parts list of both mouse and human brains.”