In biology, a mutation is the permanent alteration of the nucleotide sequence of the genome of an organism, virus, or extra chromosomal DNA or other genetic elements. A mutation is a change in DNA, the hereditary material of life. Mutation can result in many different types of change in sequences. Mutations in genes can either have no effect, alter the product of a gene, or prevent the gene from functioning properly or completely.
Mutations result from damage to DNA, which is not repaired, errors in the process of replication, or from the insertion or deletion of segments of DNA by mobile genetic elements. Mutations play a part in both normal and abnormal biological processes including evolution, cancer, and the development of the immune system, including Junctional diversity.
Previously, we shared about a ‘Revolutionary Nano-thermometer made from DNA’. Scientists made this Nano-thermometer from actual DNA. This new device is nearly about 20,000 thousand times tinier as compared to human hair. It is reversible, robust and efficient.
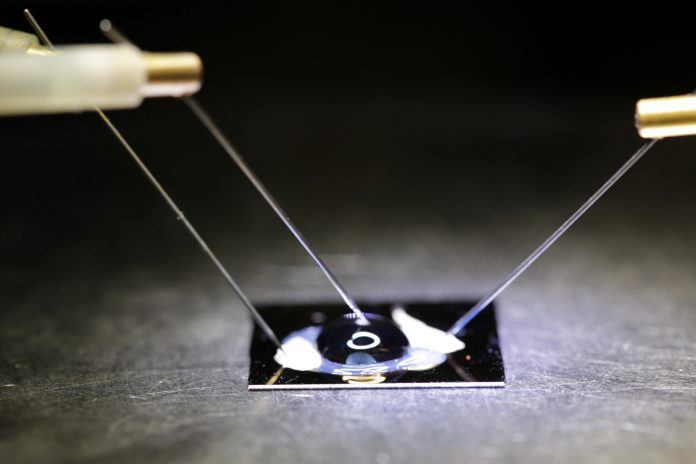
Recently, Bioengineers at the University of California, San Diego have cultivated an electrical graphene chip. This chip is developed with the purpose to check mutations in DNA. According to researchers, this technology has lots of applications in the medical field like blood-based tests for early cancer protection, controlling disease biomarkers and actual time detection of viral and microbial sequences.
This technology, which is at demonstration level, is the first step toward a biosensor chip that can be inserted into the body to check a specific DNA mutation. It gives result in actual time. It also can give the result or information wirelessly to a mobile device, smartphone or laptop.
Ratnesh Lal, professor of bioengineering, mechanical engineering and materials science in the Jacobs School of Engineering at UC San Diego, said that they are at the beginning of developing a fast and low-cost digital method to detect gene mutations at high resolution. It is on the scale of a single nucleotide change in a nucleic acid sequence.
Lal, the team leader, and Gennadi Glinsky, a research scientist at IEM, cultivates a new technique to identify the most common genetic mutation called a single nucleotide polymorphism (SNP). SNP is differentiation of a single nucleotide base (A, C, G or T) in the DNA sequence. Most of the SNPs have no recognizable effect on health. Some of them are related to pathological conditions such as cancer, diabetes, heart disease, neurodegenerative disorders, autoimmune and inflammatory diseases.
Currently, the technique used for identifying are almost slow and expensive. It needs using of heavy devices.
Preston Landon, a research scientist in Lal’s research group said that “they are developing a simple, fast, low-cost and portable way to identify SNPs by using a small chip. It can work with your cell phone too.”
How this New Biosensor Chip was made?
This chip is made from a DNA probe equipped on a graphene field effect transistor. This DNA probe is a devised piece of double fine treated DNA. It contains a sequence coding for a specific type of SNP. Along with the single nucleotide mutation, this chip is specially devised and manufactured to capture DNA or RNA molecules. Whenever the molecules of DNA or RNA constrained to the probe, an electrical signal is produced.
How this biosensor chip work?
This chip works by performing a displacement of fine treated DNA. This is the process in which a DNA double helix exchanges one fine thread for another complementary fine thread. The new complementary fine thread contains the single nucleotide mutation. It binds more strongly to one of the fine thread in the double helix and displaces the other fine thread.
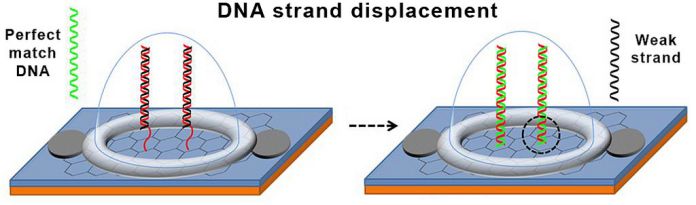
In this study, the DNA investigation is a double helix containing two complementary DNA fine threads that are devised to bind weakly to each other: the first one is a “normal” fine thread. It is attached to the graphene transistor. And another one is “weak” fine thread, in which four the G’s in the sequence were replaced with inosines to weaken its bond to the normal fine threads.
DNA fine threads have the perfectly matching complementary sequence to the normal fine thread. In another word, threads that contain the SNP that will bind to the normal threads and knock off the weak strand. Researchers devised the chip to produce an electrical signal when an SNP-containing thread binds to the probe. Through this, it allows for quick and easy SNP detection in a DNA sample.
Some key features:
- The DNA probe is attached to a graphene transistor, enables the chip to run electronically.
- It can perform DNA threads displacement on a graphene field effect transistor.
- This is the first example of combining dynamic DNA nanotechnology with high-resolution electronic sensing.
- It can be used with your wireless electronic devices to detect SNPs.
- It enables the chip to detect an SNP within longer stretches of DNA.
- This probe is 47 nucleotides long, the longest DNA probe that has been used in SNP detection
This technology overcomes the problems of current SNPs detection techniques, which commonly use single-stranded DNA probes. Through a double threaded DNA probe, only a DNA thread that’s a perfect match to the normal thread is capable of displacing the weak thread.
Lal said, ” A single threaded DNA probe doesn’t provide this selectivity even a DNA thread containing one mismatching nucleotide base can bind to the probe and generate false-positive results.”
Full paper: “Highly specific SNP detection using 2D graphene electronics and DNA strand displacement,” by Michael T. Hwang,* Preston B. Landon,* Joon Lee, Duyoung Choi, Alexander H. Mo, Gennadi Glinsky, and Ratnesh Lal, all from UC San Diego.
