Researchers at the Institute of Science and Technology Austria (IST Austria) have figured out how to control the conduct of individual bacteria by associating them to a PC. Scientists utilized the setup to construct a hereditary circuit that is mostly living and incompletely computerized.
In the test evidence of idea, they made quality articulation in microorganisms waver and controlled the examples of swaying by altering advanced correspondence between singular microscopic organisms.
Scientists wanted to design a microorganism that can satisfy a specific undertaking as a major aspect of its metabolic cycle, for example, creating a malignancy medicate or an anti-microbial, They thus need to roll out a critical number of improvements to the first life form.
Each of these progressions has a few impacts that could meddle with the impacts of every other change, modifying the last outcome.
Experimental biologist Remy Chait said, “Even if you understand what the different parts do, you don’t know what happens when you put them together. There is feedback between them that makes the behavior of the full circuit unpredictable.”
A potential solution to this problem comes from software development and is called unit and integration testing. For that purpose, scientists tested each component individually and its interaction with the surroundings. They did this by stimulating surroundings in a virtual space and to let the component interact with this virtual world. Scientists now planning to apply this method to biological systems.
Remy Chait said, “Biological systems are complex, and we would benefit if we could debug them like a computer code. In unit and integration testing, you simulate the environment and plug each of the components in separately to verify that they function as intended. Then you combine them in pairs and start all over. In this way, you will see at which point feedback and interference start to disturb the system, and adjust it appropriately.”
“By iterating this method, the virtual part could be steadily reduced until the system is fully biological again, and has the desired function.”
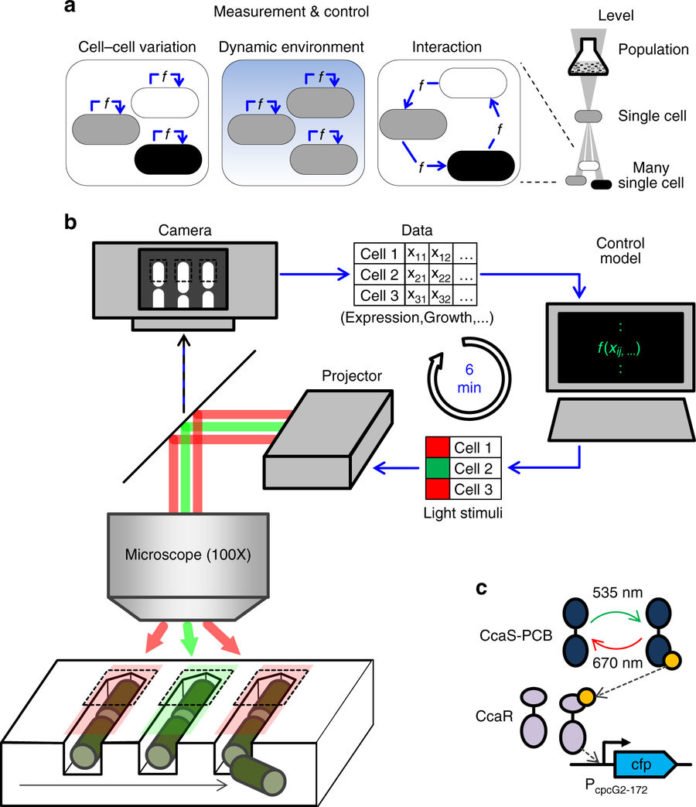
Scientists then demonstrated the feasibility of bio-digital hybrids with a bio-digital oscillator. During experiments, they modified E.coli cells to generate a protein that fluorescents blue-violet. This colored light forms the interface with the digital side.
After every 6 minutes, the PC measures how much light the phone creates, and gathers a virtual flag particle in the extent of it. At the point when the flag surpasses a specific edge, creation of the fluorescent protein by the cell is turned off.
This is finished by a projector that tasks red or green light as an “off” or “on” flag onto the light-touchy cells and along these lines interfaces the computerized segment back to the living parts of the circuit.
Remy Chait said, “The cells are interacting with the simulated environment. What they do influences what the computer does, and what the computer does influence the reaction of the cells. If you know Star Trek, you have certainly heard of the Holodeck. What we have built is essentially a simple Holodeck for genes of microorganisms.”
At the point when the specialists tried their half and half circuits, the number of inhabitants in cells shined in blue-violet—and the sparkle swayed, though with varieties between the individual microbes.
Be that as it may, scientists needed the microscopic organisms to sway in synchrony, so they changed the computerized part and set up a virtual correspondence arrange between the microorganisms. In this set-up, a portion of the virtual flag is conveyed amongst neighbors and the gathering of microbes show distinctive sorts of aggregate wavering.
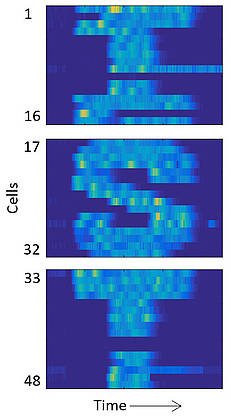
A different application of the researcher’s platform is feedback control of individual cells that guide them along pre-specified trajectories of fluorescent gene expression. In this way, they could make a group of cells trace pictures or letters over time.