In a new study, scientists at the Brown University portrayed the detailed structure of proteins that empower sperm-egg fusion in two unique species: a flowering plant and a protozoan. The specialists believe that noteworthy the procedure in these species and their relatives may convey researchers a bit nearer to understanding it crosswise over sexual species, including humans and different vertebrates.
Sperm-egg fusion is the very primary step that leads to new individuals in sexually reproducing species. Although, scientists are now examining the complexities of how it works.
Mark Johnson, an associate professor of biology at Brown University and a study co-author said, “It’s surprising to me that we still don’t know how a human sperm fuses with a human egg. One of the things we hope this paper will do is establish a structural signature for the proteins that make gamete fusion work in these species so that we might be able to look for it in species where those protein mechanisms are still unknown.
In the mid-2000s, he distinguished a protein on the sperm membrane of the blooming plant species Arabidopsis thaliana that appeared to have some impact on the gamete fusion process in that species. His work demonstrated that a change in the quality the produces the protein, known as HAP2, causes sperm to wind up unable to meld with the Arabidopsis egg.
After some years, Kristin Beale, a Brown graduate student working with Johnson, predicted that HAP2 was related to the protein that viruses use to bore their way into host cells.
Johnson said, “That makes sense since both sperm and viruses need to have a mechanism for inserting themselves into a cellular membrane.”
Since discoveries, scientists have searched the genomes of other organisms for gene sequences that look similar to the Arabidopsis HAP2. They’ve discovered comparable groupings in a wide assortment of eukaryotes (creatures whose cells have a discrete core) — many plant species, green growth, bugs and a few creatures. It was prominently absent however in vertebrates, including humans.
Johnson said, “It’s possible, that vertebrates do indeed have a HAP2-like protein, but its genetic sequence may have changed so much over evolutionary history that it’s hard for researchers to spot by sequence alone. So rather than searching for genes that produce the protein, it might be better to look at the structure of proteins themselves — to search for vertebrate proteins that are structurally similar to those of the HAP2 proteins that have already been identified. But that requires the structures of known HAP2 proteins to be resolved in detail, which is what Johnson and his colleagues set out to do with this latest study.”
Scientists then resolved the structures of HAP2 proteins from two distantly related eukaryote species: Arabidopsis and Trypanosoma cruzi, a protozoan parasite. For this, the used a technique called x-ray crystallography, which involves crystallizing the proteins and then observing how the crystals scatter x-rays.
The structure of the protein can be seen from the scrambling design. Scientists spend significant time in the strategy, especially to image the viral fusion proteins that are identified with HAP2. In addition, they determined that how HAP2 drives fertilization in flowering plants, defining the amino acids at the tip of the protein that inserts into the egg membrane.
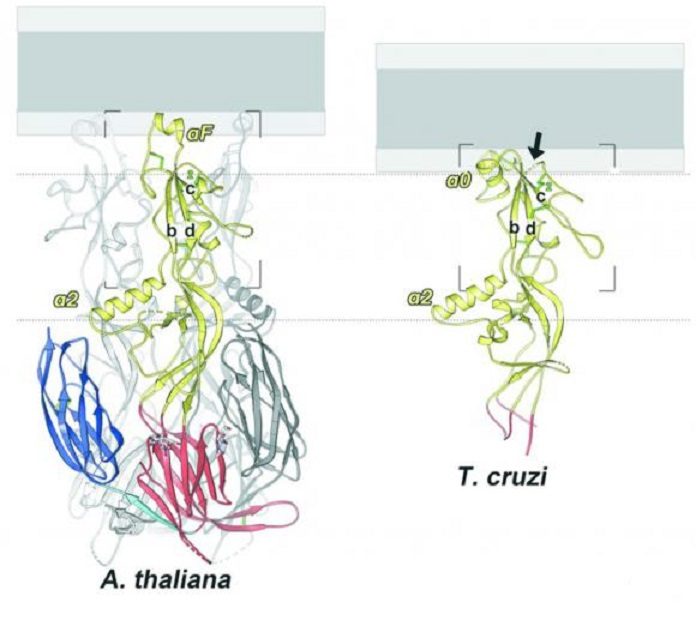
The examination found that while the fundamental structure of the HAP2 proteins from the two species was extensively comparative, they had developed to be diverse in key areas. In particular, the tips of the proteins — the parts that are thought to first pierce the film of an egg cell — were significantly unique.
While the streaming plant HAP2 utilizes a solitary helical structure at the tip, the protozoan variant has three little circles that guide into the objective layer. While the distinctions in addition components between the two proteins shed new light on how HAP2 functions, the likenesses between the two are imperative in searching for HAP2 in different life forms.
Johnson said, “At this point, we have two kinds of organisms: those for which we know how gamete fusion works — and they all have HAP2 — and organisms for which we don’t know how gamete fusion works, none of which have a recognizable HAP2. Now that we know what’s conserved structurally in these two proteins that differ at the sequence level, we can think about looking for those structures to look for related proteins in other species.”
“That includes humans. We just need to get more structures for human proteins so we can look for common structural features.”
Scientists published their paper in the journal PLOS Biology. Other authors on the paper were Pierre Legrand, Gérard Péhau-Arnaudet, Ahmed Haouz and Thomas Krey.