By using light and genetically engineered bacteria, Stanford scientists have shaped the growth of bacterial communities. They have developed a technique that renders intricate designs of polka dots to stripes to circuits overnight.
The technique could potentially grow sticky films of bacteria into elaborate microscopic images. It could expose how potentially dangerous biofilms grow and transmit antibiotic resistance and thus lead to novel biomaterials or synthetic microbial communities.
Ingmar Riedel-Kruse, assistant professor of bioengineering and senior author of the paper said, “Most of the bacteria on Earth live in biofilm communities and biofilms are very relevant in disease in health – plaque on our teeth or catheter-based bacterial infection, for example. Understanding how biofilms function is an important question on many levels.”
“The technique could clarify how biofilms grow and lead to the development of novel biomaterials or synthetic microbial communities that could be implemented in small devices or systems, such as microfluidic chips or biofilm-based circuits.”
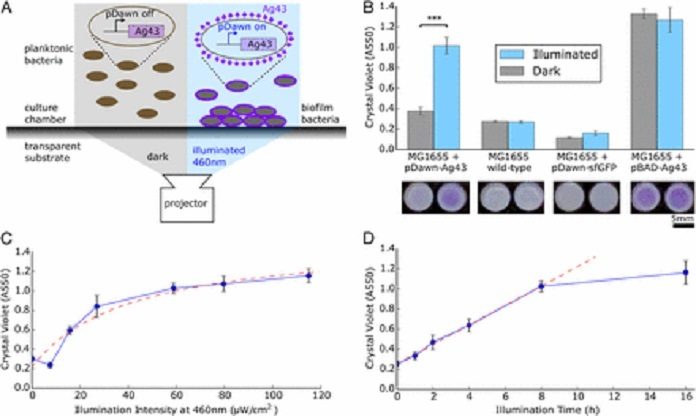
The technique is basically based on E. coli bacteria to discharge a sticky protein because of a specific wavelength of blue light. When they sparkle the fitting wavelength light in the coveted example on a culture dish of altered microorganisms, the microscopic organisms adhere to the lit zones, framing a biofilm in the state of the example. The specialists call their strategy biofilm lithography for its closeness to lithography utilized as a part of making electronic circuits.
Different systems for designing bacterial groups exist, incorporating keeping them with an inkjet printer or pre-designing the way of life surface with chemicals that predisposition bacterial development in particular zones. Be that as it may, biofilm lithography has the advantage of speed, straightforwardness, higher determination, and similarity with an assortment of surface conditions including shut microfluidic gadgets, the scientists said.
The perplexing outlines made conceivable with biofilm lithography could help in investigating the elements of bacterial groups.
Xiaofan Jin, a graduate student in bioengineering and lead author of the paper said, “Biofilms exist in a social environment with other bacteria. Interactions between these bacteria are often dictated by where they grow relative to each other and this could be a great tool for specifying exactly when and where in a bacterial community certain species can live.”
Riedel-Kruse said, “In the literature, there are different models of how certain bacterial species form biofilms. We argue, at least with this species, that we provided additional evidence for that one hypothesis.”
By coincidence, the 25-micrometer resolution the researchers achieved with biofilms is similar to the first silicon photolithography, which contributed to the widespread success of silicon semiconductors. Similarly, the researchers see many versatile and impactful applications for their bacterial designs.
“We’re hoping this tool can be applied toward further understanding of bacterial communities, both natural and synthetic,” said Jin. “We also see potential in having these communities do useful things, such as metabolic biosynthesis or distributed biocomputation. It may even be possible to create novel biomaterials such as conductive biofilm circuits.”
The technique, described in the March 19 Proceedings of the National Academy of Sciences.