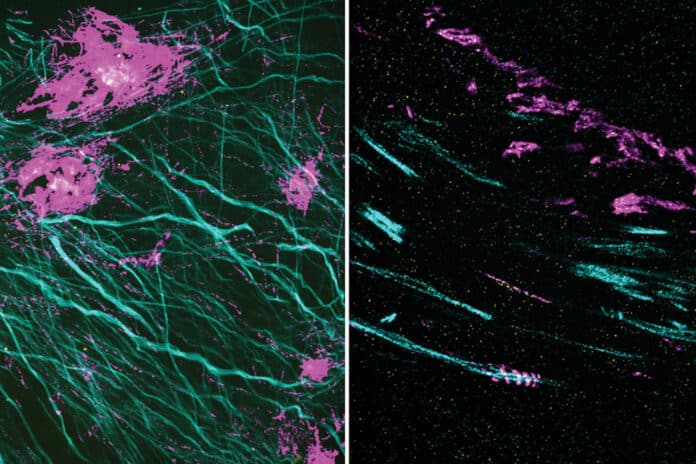
Many crowded biomolecular structures in cells and tissues are inaccessible to labeling antibodies. Understanding how proteins within these structures are arranged with nanoscale precision requires that these structures be decrowded before labeling.
Sometimes these structures can be difficult to image because the fluorescent labels used to make them visible can’t wedge themselves in between the molecules. To overcome this limitation, MIT scientists have developed a new technique that makes invisible molecules visible. The method helps scientists de-crowd the molecules by expanding a cell or tissue sample before labeling the molecules. This allows the molecules to be more accessible to fluorescent tags.
Edward Boyden, the Y. Eva Tan Professor in Neurotechnology, a professor of biological engineering and brain and cognitive sciences at MIT, said, “It’s becoming clear that the expansion process will reveal many new biological discoveries. Suppose biologists and clinicians have been studying a protein in the brain or another biological specimen, and they’re labeling it the regular way. In that case, they might be missing entire categories of phenomena.”
As scientists noted, this method allows scientists to visualize molecules and cellular structures that have never been seen before. In their study, scientists could image a nanostructure found in the synapses of neurons. They also imaged the structure of Alzheimer’s-linked amyloid beta plaques in greater detail than possible.
Deblina Sarkar, an assistant professor in the Media Lab and one of the study’s lead authors, said, “Our technology, which we named expansion revealing, enables visualization of these nanostructures, which previously remained hidden, using hardware easily available in academic labs.”
Before expanding the tissue, scientists in the original version of expansion microscopy applied fluorescent markers to molecules of interest. The labeling was carried out first, partly because the sample’s proteins needed to be broken down by an enzyme before the tissue could be enlarged. This meant that once the tissue was stretched, it was impossible to mark the proteins.
To overcome that issue, scientists ad to find a way to expand the tissue while leaving the proteins intact. Instead of enzymes, they used heat to soften the tissue. This enabled the tissue to expand 20-fold without being destroyed. They then separated proteins that could be labeled with fluorescent tags after expansion.
As scientists could access numerous proteins for labeling, they could identify tiny cellular structures within synapses. These nanocolumns are believed to help make synaptic communication more efficient.
Jinyoung Kang, an MIT postdoc, said, “This technology can be used to answer many biological questions about dysfunction in synaptic proteins, which are involved in neurodegenerative diseases. There has been no tool to visualize synapses very well.”
Using their new technique, scientists imaged beta-amyloid, the plaque-forming peptide in AD. They used brain tissue from mice and found that amyloid beta forms periodic nanoclusters. This was not been seen before.
Surprisingly, they also found potassium channels in the amyloid beta clusters. Plus, the amyloid beta molecules generate helical structures along axons.
Margaret Schroeder, an MIT graduate student who is also the author of the paper, said, “In this paper, we won’t speculate as to what that biology might mean, but we show that it exists. That is just one example of the new patterns we can see.”
Sarkar said, “I’m fascinated by the nanoscale biomolecular patterns this technology unveils. With a background in nanoelectronics, I have developed electronic chips that require exact alignment in the nano fab. But when I see that Mother Nature has arranged biomolecules with such nanoscale precision in our brain, that blows my mind.”
Journal Reference:
- Sarkar, D., Kang, J., Wassie, A.T. et al. Revealing nanostructures in brain tissue via protein decrowding by iterative expansion microscopy. Nat. Biomed. Eng (2022). DOI: 10.1038/s41551-022-00912-3