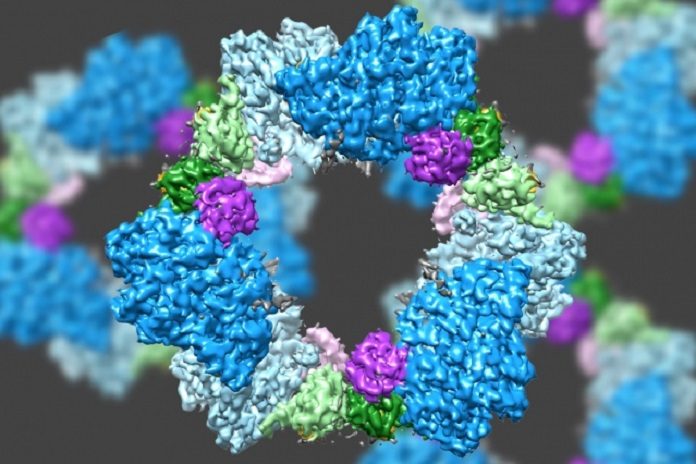
MIT scientists have investigated the enzyme’s structure by using a technique called electron microscopy. This new discovery also unveiled the likely mechanism for how cells regulate the enzyme, known as ribonucleotide reductase (RNR).
Enzyme structure is crucial for maintaining an adequate supply of DNA building blocks in human cells. According to scientists, the technique displayed the mechanism in such way that it could be used to design antibiotics that selectively block the bacterial enzyme.
Catherine Drennan, an MIT professor of chemistry and biology said, “People have been trying to figure out whether there is something different enough that you could inhibit bacterial enzymes and not the human version. By considering these key enzymes and figuring out what are the differences and similarities, we can see if there’s anything in the bacterial enzyme that could be targeted with small-molecule drugs.”
The RNR compound, which is found in every single living cell, changes over ribonucleotides (the building squares of RNA) to deoxyribonucleotides (the building pieces of DNA). Cells must keep an adequate store of these building squares, however, when they amass too much, RNR is stopped by a deoxynucleotide particle known as dATP. At the point when more deoxynucleotides are required, a related atom called ATP ties to RNR and walks out on.
An unordinary highlight of RNR is that it can catalyze the creation of four unique items: the nucleotide bases frequently abridged as A, G, C, and T. Scientists had previously discovered that the enzyme achieves this by changing its shape in response to regulatory molecules.
Previous studies on RNR structure focused on the version found in E. coli. Those studies used X-ray crystallography that reveals the atomic and molecular structure of a protein after it has been crystallized.
In this new study, scientists examined the human version of RNR, that turned out to be very different from the bacterial version, proved elusive using X-ray crystallography, which doesn’t work well for proteins that don’t readily crystallize. Thus, scientists used an advanced form of microscopy known as cryo-electron microscopy (cryo-EM).
cryo-EM normally offered a determination of around 10 to 20 angstroms, which may uncover the general state of a protein, however, no insight about the position and state of littler basic units inside it.
Scientists already had an idea that the RNR composed of two proteins subunits known as alpha and beta. The technique reveals that the human variant of the compound structures a ring produced using six of the alpha subunits. Whenever ATP, which actuates RNR, is bound to the protein, the ring is unsteady and can be effortlessly opened up, enabling the beta subunit to advance into the ring.
This joining of alpha and beta permits the protein’s dynamic site, situated in the beta subunit, to play out the substance responses important to create deoxynucleotides. Be that as it may, when the inhibitor dATP is available, the ring turns out to be substantially more unbending and does not enable the beta subunit to enter. This keeps the chemical from catalyzing the creation of deoxynucleotides.
A few tumor tranquilizes now being used or being developed focus on the human adaptation of RNR, meddling with growth cells’ capacity to repeat by restricting their supply of DNA building squares. The MIT group has discovered proof that no less than one of these medications, clofarabine diphosphate, works by actuating the arrangement of inflexible 6-unit alpha rings.
This 6-unit ring isn’t found in the bacterial type of RNR, which rather amasses into an unmistakable ring containing four alpha subunits and four beta subunits. This implies it could be conceivable to plan anti-infection agents that objective the bacterial form yet not the human adaptation.
Drennan said, “The technological advances that have allowed cryo-EM to get to such high resolution are really exciting. It’s really starting to revolutionize the study of biology.”