Antibiotics are used to kill bacteria. With time, some of these bacterias adapt to these medicines, which may change so that antibiotics can′t kill them. This is known as antibiotic resistance.
Some general antibiotic resistance causes include overprescription of antibiotics, poor hygiene, and the spread of resistant bacteria. Antibiotic use is the leading cause of antibiotic resistance.
Why is antibiotic resistance a problem?
Antimicrobial resistance in Gram-negative bacteria is one of the greatest threats to global health. It leads to higher medical costs, prolonged hospital stays, and increased mortality.
New strategies for fighting antibiotic resistance are, therefore, urgently required.
Scientists, including experts from Imperial College London, found a new approach to fighting antibiotic resistance that causes human disease, such as E. coli, K. pneumonia, and P. aeruginosa. Their impairing antibiotic-resistant bacteria represents an entirely new way of thinking about targeting resistance.
Bacterias in AMR host several proteins in their arsenals that neutralize antibiotics. To function properly, these resistance proteins need to be folded in the right shapes.
A protein called DsbA helps fold resistance proteins into suitable shapes to neutralize antibiotics.
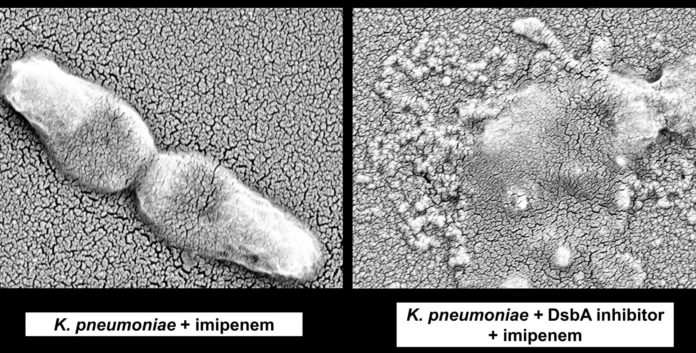
For their study, scientists used chemicals to inhibit DsbA. These chemicals can not be used directly in human patients.
Dr. Chris Furniss, one of the lead authors for the Department of Life Sciences study at Imperial, said: “Since the discovery of new antibiotics is challenging, it is crucial to develop ways to prolong the lifespan of existing antimicrobials.”
“Our findings show that by targeting disulfide bond formation and protein folding, it is possible to reverse antibiotic resistance across several major pathogens and resistance mechanisms.”
“This means that the development of clinically useful DsbA inhibitors in the future could offer a new way to treat resistant infections using currently available antibiotics.”
Scientists hope to ultimately combine a DsbA inhibitor with existing antibiotics to restore the drugs’ ability to kill bacteria.
Nikol Kaderábková, previously a graduate student at Imperial and currently a postdoctoral researcher at UT Austin, and the second lead author of the study, said: “We reasoned that if DsbA is required for the folding of resistance proteins, preventing it from working would indirectly inhibit their function.”
Journal Reference:
- R Christopher D Furnis et al. Breaking antimicrobial resistance by disrupting extracytoplasmic protein folding. DOI: 10.7554/eLife.57974