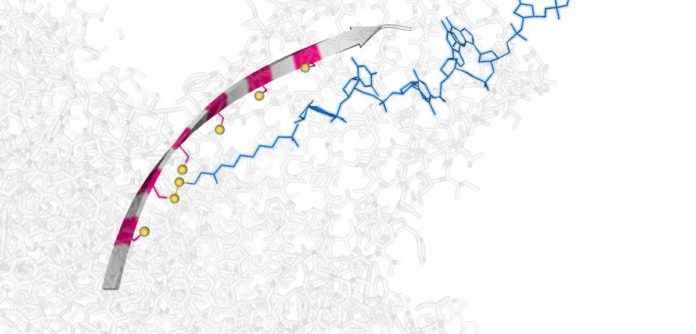
Scientists at the University of Oxford have developed a “molecular hopper” that can move single strands of DNA through a protein nanotube. This hopper works by making and breaking in sequence basic chemical bonds that join it to a nanoscale track.
Moreover, the user can turn it on or off using a small electrical potential, which ultimately might make it suitable for use in nanopore DNA sequencing devices.
Professor Hagan Bayley of Oxford University’s Department of Chemistry, who led the research, said: “Being able to control molecular motion is the holy grail of building nanoscale machines. Being able to process single molecules of DNA under precise chemical control may provide an alternative to the use of enzymes in DNA sequencing technologies, improving their speed and the number of molecules that can be analyzed in parallel.”
The hopper currently takes a couple of moments for each progression, and the specialists currently looking to increase the speed of the science and in addition the length of the track, which is by and by constrained to six decent footings.
The hopping motion utilizes extremely simple chemistry in view of 3 sulfur particles [thiol/disulfide interchange], which happens in water at room temperature. The hopper makes sub-nanometer strides (0.7 nm), and is fueled and controlled by an electric field; the heading of hopping can be exchanged by switching the electric field. This is observed progressively at the single particle level.
A tightening movement is required for nanopore sequencing, which at the display is accomplished by utilizing an enzyme. The hopping movement in the recently distributed device is a chemical ratchet and this rule may be connected to DNA and RNA sequencing on the grounds that the progression estimate is like the between nucleotide distance in single-stranded DNA.
Scientists have published their research in the journal Science.