A team of researchers from Stanford University in Palo Alto, California and University of California, Los Angeles (UCLA) have developed a third approach based on a technique called electron diffraction to determine the precise shape of small organic molecules. It is unlike the other two techniques utilized before.
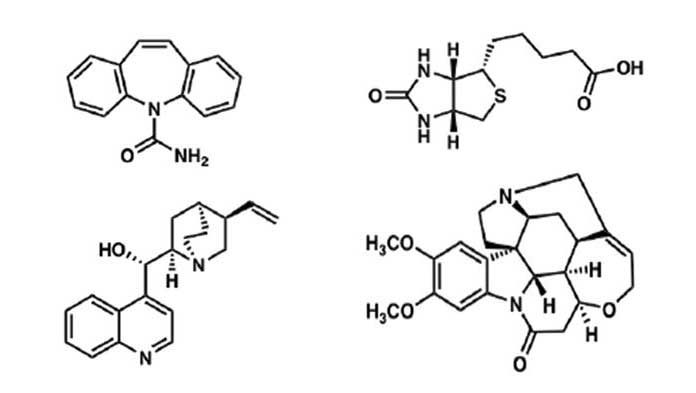
In chemistry, structure rules since it decides how a molecule behaves. But the two ideal methods to map the structure of small organic molecules, such as pharmaceuticals, hormones, and vitamins, have flaws. The novel discovered approach works well with almost to become invisible samples and is surprisingly easy.
Carolyn Bertozzi, a chemist at Stanford University in Palo Alto, California, said, “I am blown away by this.”
“The fact that you can get these structures from [a sample] a million times smaller than a speck of dust, that’s beautiful. It’s a new day for chemistry.”
One of the two methods utilized to determine chemical structures earlier was x-ray crystallography. A beam of x-rays is fired at a pure crystal containing millions of copies of a molecule lined up in a single orientation. Later the researchers track the x-rays bounce off atoms in the crystal to determine the position of every atom in the molecule.
Crystallography can pinpoint atomic positions down to less than 0.1 nanometers, about the size of a sulfur atom. Yet, the technique works best with fairly large crystals, which can be hard to make.
Brian Stoltz, an organic chemist at the California Institute of Technology (Caltech) in Pasadena, said, “The real lag time is just getting a crystal.”
“That can take weeks to months to years.”
The second technique, investigated as nuclear magnetic resonance (NMR) spectroscopy, doesn’t require crystals. It concludes structures by perturbing the magnetic behavior of atoms in molecules and then tracking their behavior, which changes based on their atomic neighbors. Besides, NMR requires a fair amount of starting material. Whereas this method is implicit and can lead to mapping flaws with larger druglike molecules.
How does the novel approach work? It is based on a technique called electron diffraction, which discharges an electron beam through a crystal and, as in x-ray crystallography, determines structure from diffraction patterns.
Specifically, this approach has been found helpful in solving the structure of a class of proteins lodged in cell membranes. A team first form tiny 2D sheetlike crystals of multiple copies of a protein wedged in a membrane in this particular instance.
But in other instances endeavors to grow the protein crystals go wrong. What happened in those instances is that despite getting single-membrane sheets, researchers end up with numerous sheets pile up one another, which can’t be analyzed by conventional electron diffraction. And the crystals can be too compact for x-ray diffraction.
Tamir Gonen, an electron crystallography expert at the University of California, Los Angeles (UCLA), said, “We didn’t know what to do with all these crystals.”
Eventually, his team varied the technique. This time despite firing their electron beam from one direction at a static crystal, they rotated the crystal and tracked how the diffraction pattern changed. Also, instead of a single image, they produced something more like molecular computerized tomography scan. That entitled them to get structures from crystals one-billionth the size of those required for x-ray crystallography.
Bertozzi stated, “I’ve had dreams in my life where I’m looking through a microscope and I see a molecular model with balls and sticks.”
“They basically find some microcrystalline schmutz on an EM [sample holder], take some data, and there are the balls and sticks I dreamed about. It’s unbelievable it works so well.”
Bertozzi and others said, “Because it does work so smoothly, the new technique could revolutionize fields both inside and outside of research.”
But only about one-quarter to one-third of the compounds form crystals big enough for x-ray crystallography.
Tim Grüne, an electron diffraction expert at the Paul Scherrer Institute in Villigen, Switzerland, said, “This will remove a bottleneck and lead to an explosion of structures.”
This new approach could help to speed up the search for promising drug leads in tiny samples of exotic plants and fungi. For crime labs, it could be a blessing since it will help them promptly determine the latest heroin derivatives hitting the streets.
It could also help for the Olympics officials clean up sports by making it easier to spot vanishingly small amounts of performance-enhancing drugs. All because structures rule—and are now easier than ever to decipher.
This research published in the journal Science.