Hepatitis B is the most common form of hepatitis. Usually transmitted through blood or sexual relations, HBV is responsible for one of the most severe and common infectious diseases.
This virus infects the liver cells, resulting in a brief inflammation of the liver that may progress to a chronic infection. Severe diseases like cirrhosis or liver cancer can result from this. There is no effective treatment for the chronic form of the disease, which can only be prevented by vaccination.
Scientists from the University of Geneva identified an essential protein complex that is active when the virus infects our body. They deciphered the precise functioning of this protective mechanism.
A UNIGE team led by Michel Strubin, an associate professor in the Department of Microbiology and Molecular Medicine and the Geneva Centre for Inflammation Research at the UNIGE Faculty of Medicine, discovered a mechanism for this disease in 2016 that is essential for understanding: when our immune system defends itself against it, a complex, or an interdependent set, of six proteins called SMC5/6, present in our cells, detects the viral DNA and blocks it.
The virus then responds by releasing a unique protein called the X protein. This protein penetrates the cell and breaks down SMC5/6, rendering it incapable of performing its sentinel role.
SMC5/6’s role in antiviral defense was unknown before this finding. It was discovered to be crucial for keeping our chromosomes’ structural integrity. A new development has been made by Michel Strubin’s team today. The UNIGE researchers have determined the three processes and the specific proteins needed for SMC5/6 to perform its antiviral function in a study that was carried out in partnership with the American pharmaceutical company Gilead Sciences.
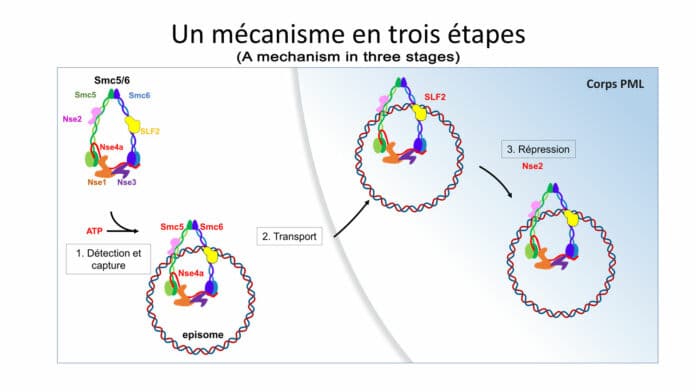
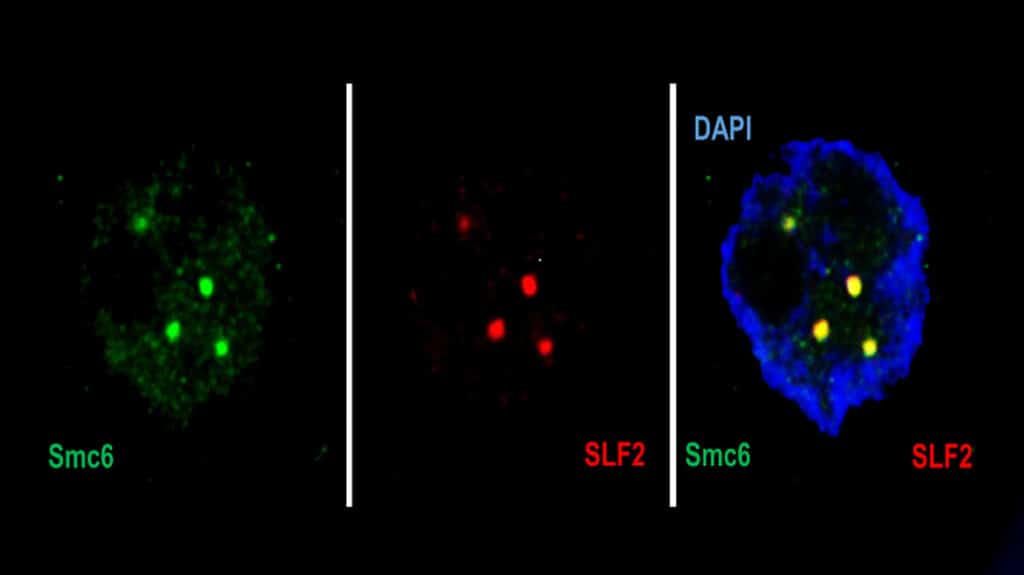
Fabien Abdul, a senior research and teaching assistant in the Department of Microbiology and Molecular Medicine at the UNIGE Faculty of Medicine, said, “In the first step, a protein of the SMC5/6 complex detects the virus’ DNA and traps it. Then, a second protein of the complex – SLF2 – takes the trapped DNA of the virus into a sub-compartment of the nucleus of the attacked cell, called the PML body. A third protein – Nse2 – then comes into play and inhibits the virus’ chromosome.”
“As SMC is a large family of protein complexes, the researchers also wanted to know whether other ‘members’ of this family were able to bind to hepatitis B viral DNA. We discovered that this competence was unique to SMC5/6.”
Michel Strubin, the study’s last author, said, “To achieve these results, we worked on in vitro cell cultures. We used molecular biology techniques and, more specifically, genetic scissors called CRISPR-Cas9. This tool allowed us to cut the DNA strands within the cells and thus delete or modify the gene coding for each protein constituting the SMC5/6 complex.”
“Thanks to this technique, we could make one or other of the proteins disappear and thus understand their respective functions within the complex. Based on these observations, the three steps of the antiviral mechanism could be established.”
Aurélie Diman, a postdoctoral researcher in Michel Strubin’s laboratory, said, “This discovery provides a better understanding of how the complex functions during its antiviral action. It could thus pave the way for identifying new therapeutic targets to combat the hepatitis B virus. The next stage of research will consist of better deciphering the virus’s mechanism of inhibition in the cell nucleus’s sub-compartment.”
Journal Reference:
- Abdul, F., Diman, A., Baechler, B. et al. Smc5/6 silences episomal transcription by a three-step function. Nat Struct Mol Biol 29, 922–931 (2022). DOI: 10.1038/s41594-022-00829-0