Scientists at the Eawag, the ETH and the EPFL have developed a computer model that simulates where and how often the species are present in bodies of water.
Luca Carraro, an Eawag researcher said, “If one now takes water samples and decodes the environmental DNA (also known as eDNA) therein, one knows which species cavort in which waters. One thus discovers rare species that literally fall through the net during normal testing. To be sure, this concept of eDNA has long been known. But: Until now one could determine from eDNA whether a species is present or not. But how this species is distributed in the whole ecosystem was unknown.”
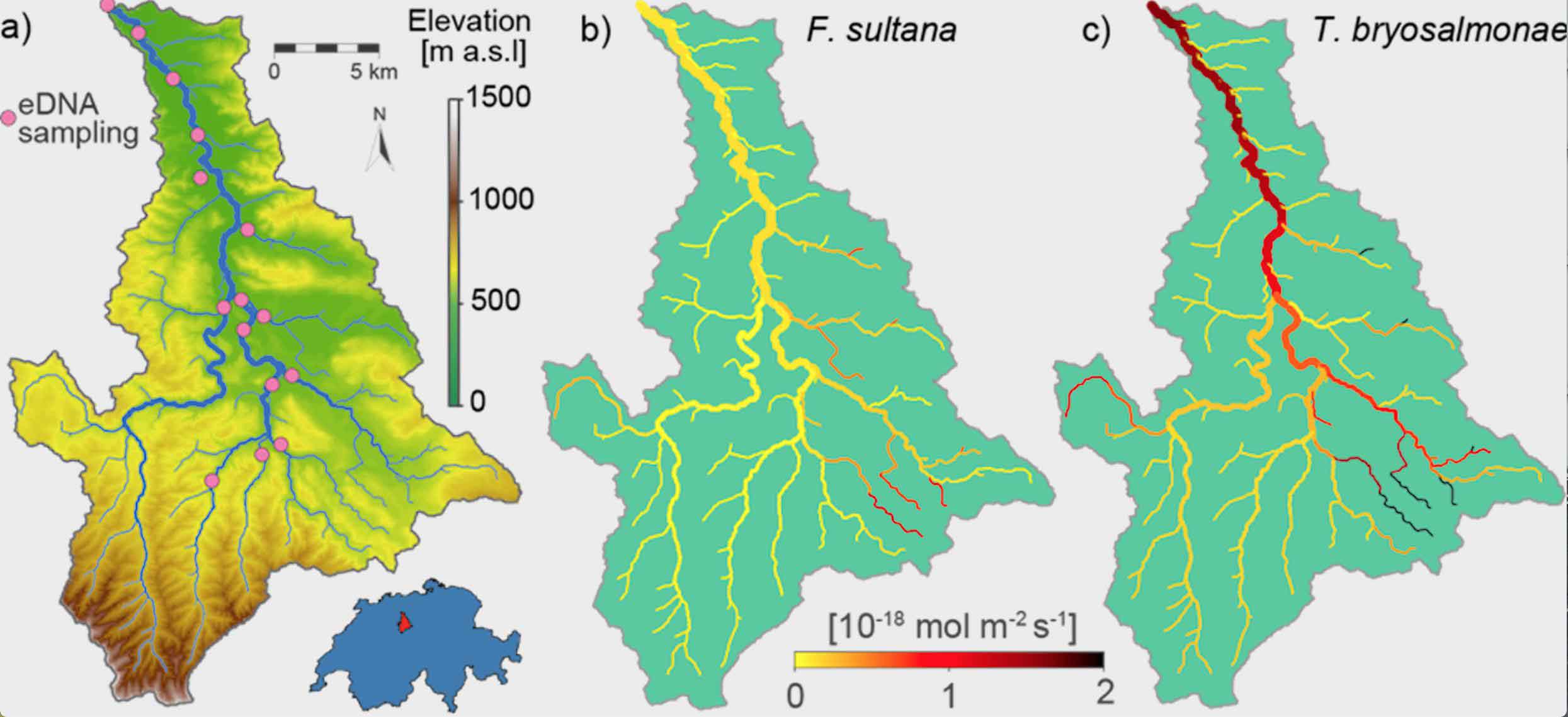
Along with scientists at the ETH and the EPFL, scientists have reconstructed the extensive presence of species from selective eDNA measurements. They used the method to determine the biomass distribution of a bryozoan (Fredericella sultana) and its parasite (Tetracapsuloides bryosalmonae) in the Wigger – a prealpine Swiss river.
The parasite causes the dreaded fish disease PKD, which carries off thousands of trout every year in Switzerland.
The computer model they developed help them determine as exactly as possible the distribution of the organisms. To make this model work, scientists fed the model with genetic material measurements.
Co-author and Eawag research group leader Hanna Hartikainen said, “This method will make it possible to monitor biodiversity all over the world simply and inexpensively. This was not possible up to now with conventional methods.”
The study is published in the journal PNAs.