People can convey the infection for quite a long time without symptoms, and 90 percent of people might be unaware that they are conveying it by any means.
However, an expected 5-10 percent of those infected may go-ahead to develop an aggressive type of leukemia or a dynamic paralytic disease.
The latest study by the Imperial College London suggests that the human leukemia virus (HTLV-1) acts at a large number of sites across the human genome, disrupting the regulation of tens of thousands of genes.
Scientists observed that how HTLV-1 interacts with human DNA when it infects a host, focusing on its target: specialized white blood cells called T-cells. They found that the HTLV-1 changes the folding pattern of human DNA in infected cells. They explain that the resulting disruption of gene function builds the risk of leukemia.
Every human cell contains around two meters of DNA, conveniently bundled into the nucleus. With a specific end goal to fit, these bending strands are firmly wound and then folded over proteins, making a thickly packed genetic spaghetti structure called chromatin.
The chromatin isn’t randomly composed, yet is collapsed into a huge number of loops, which stick out from the main ‘strand’.
These loops uncover regions of DNA to the cell’s machinery that reads and copies DNA, enabling specific chunks of the genome to be all the more easily read and transcribed.
Ongoing exploration has demonstrated that disrupting these existing loops, or making new loops, modifies the control of gene expression and might be connected to a host of diseases.
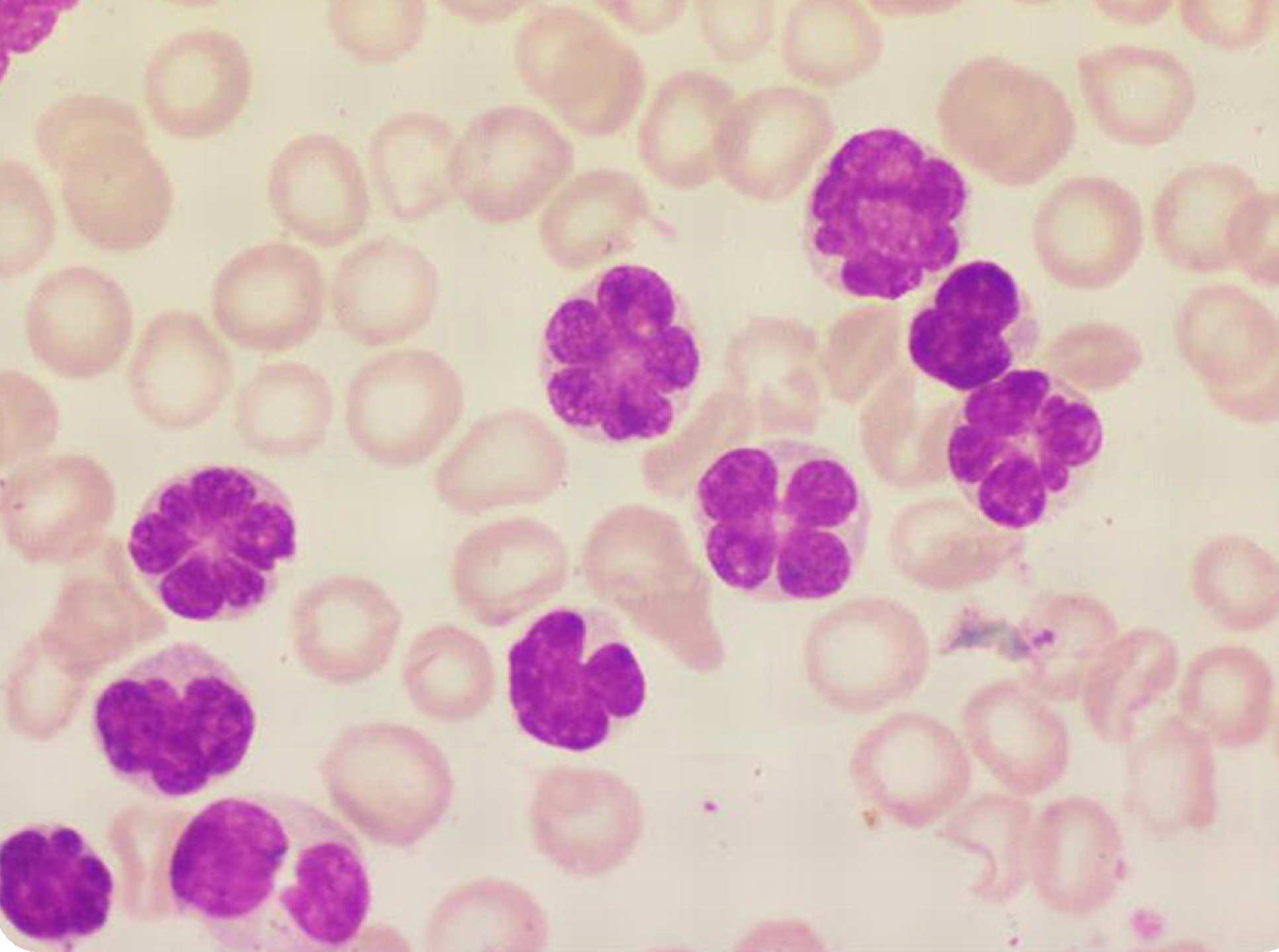
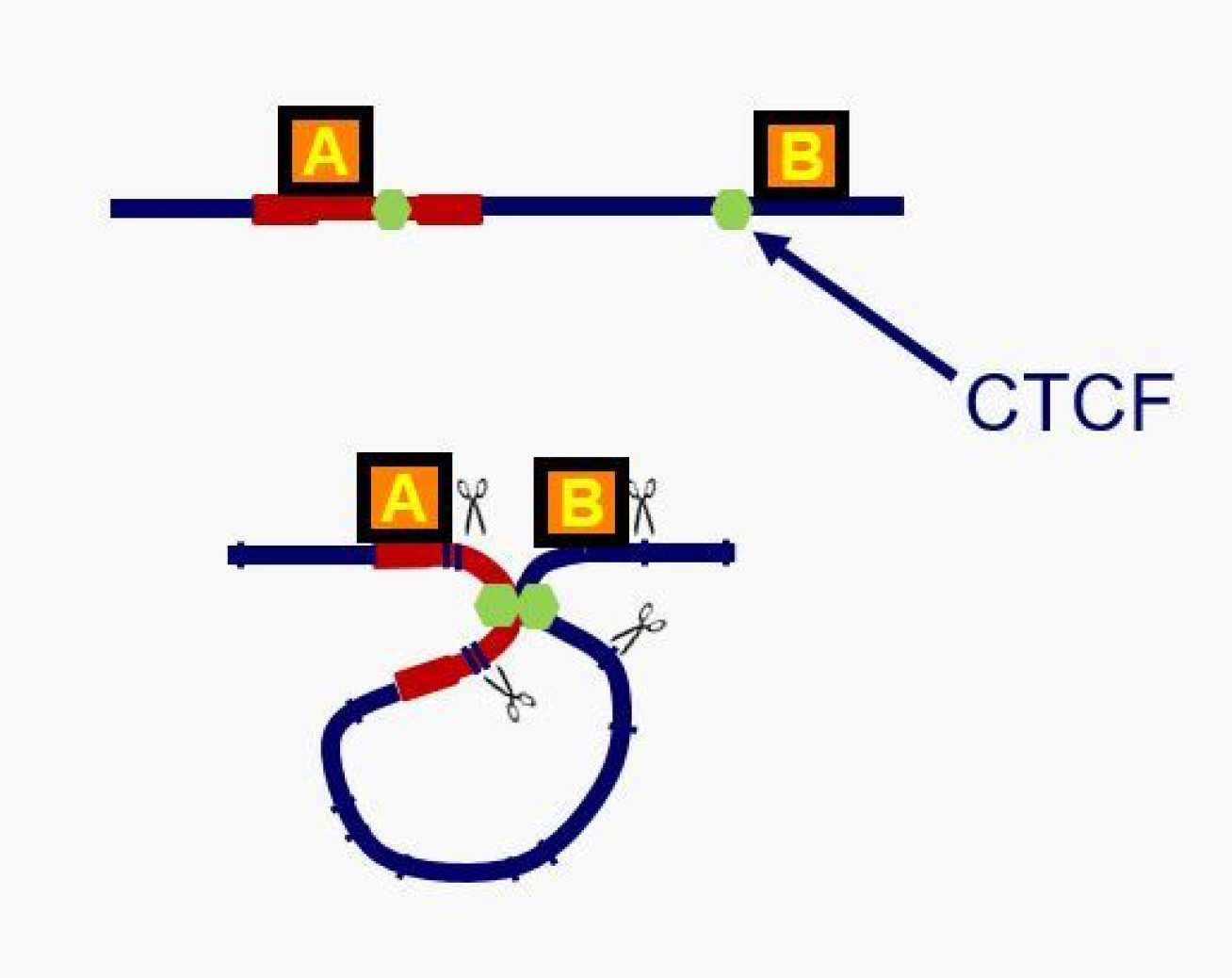
Scientists confined T-cells from HTLV-1-infected patients and analyzed which regions of the DNA were adjusted. They found that the virus binds to a protein called CTCF, which is the key protein that forms normal loops in the human genome. As a result, the virus changes the structure of the loop and the activity of genes within it.
Professor Charles Bangham, Chair of immunology in the Department of Medicine at Imperial College London and lead author of the study, said: “Through binding to these specific sites in the genome, retroviruses like HTLV-1 can alter chromatin loops and disrupt how a number of important genes are regulated. This can lead to the abnormalities and disease, such as leukemia associated with HTLV-1.”
This study offers new insight into how viruses like HTLV-1 can alter the structure of the human genome, which can result in diseases such as cancer.
Ewan Birney, Director of the European Bioinformatics Institute (EMBL-EBI) and co-author, added: “This study illustrates that by combining wet lab and dry lab expertise, we can explore previously inaccessible biological processes.
“By integrating complex genomic data – in this case, haplotype-resolved data – into our analysis, we have gained new insights into how the human T-cell leukemia virus works. In turn, this could help us understand why certain patients experience such devastating symptoms, while others are asymptomatic.”
Scientists published their research in the journal eLife.